espressomd package¶
Subpackages¶
Submodules¶
espressomd.accumulators module¶
- class espressomd.accumulators.AutoUpdateAccumulators(**kwargs)[source]¶
Bases:
ScriptObjectListClass for handling the auto-update of accumulators used by
espressomd.system.System.
- class espressomd.accumulators.Correlator(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperCalculates the correlation of two observables \(A\) and \(B\), or of one observable against itself (i.e. \(B = A\)). The correlation can be compressed using the multiple tau correlation algorithm.
The operation that is performed on \(A(t)\) and \(B(t+\tau)\) to obtain \(C(\tau)\) depends on the
corr_operationargument:"scalar_product": Scalar product of \(A\) and \(B\), i.e., \(C=\sum\limits_{i} A_i B_i\)"componentwise_product": Componentwise product of \(A\) and \(B\), i.e., \(C_i = A_i B_i\)"square_distance_componentwise": Each component of the correlation vector is the square of the difference between the corresponding components of the observables, i.e., \(C_i = (A_i-B_i)^2\). Example: when \(A\) isespressomd.observables.ParticlePositions, it produces the mean square displacement (for each component separately)."tensor_product": Tensor product of \(A\) and \(B\), i.e., \(C_{i \cdot l_B + j} = A_i B_j\) with \(l_B\) the length of \(B\)."fcs_acf": Fluorescence Correlation Spectroscopy (FCS) autocorrelation function, i.e.,\[G_i(\tau) = \frac{1}{N} \left< \exp \left( - \frac{\Delta x_i^2(\tau)}{w_x^2} - \frac{\Delta y_i^2(\tau)}{w_y^2} - \frac{\Delta z_i^2(\tau)}{w_z^2} \right) \right>\]where \(N\) is the average number of fluorophores in the illumination area,
\[\Delta x_i^2(\tau) = \left( x_i(0) - x_i(\tau) \right)^2\]is the square displacement of particle \(i\) in the \(x\) direction, and \(w_x\) is the beam waist of the intensity profile of the exciting laser beam,
\[W(x,y,z) = I_0 \exp \left( - \frac{2x^2}{w_x^2} - \frac{2y^2}{w_y^2} - \frac{2z^2}{w_z^2} \right).\]The values of \(w_x\), \(w_y\), and \(w_z\) are passed to the correlator as
args. The correlator calculates\[C_i(\tau) = \exp \left( - \frac{\Delta x_i^2(\tau)}{w_x^2} - \frac{\Delta y_i^2(\tau)}{w_y^2} - \frac{\Delta z_i^2(\tau)}{w_z^2} \right)\]Per each 3 dimensions of the observable, one dimension of the correlation output is produced. If
"fcs_acf"is used with other observables thanespressomd.observables.ParticlePositions, the physical meaning of the result is unclear.The above equations are a generalization of the formula presented by Höfling et al. [Höfling et al., 2011]. For more information, see references therein.
- Parameters:
obs1 (
espressomd.observables.Observable) – The observable \(A\) to be correlated with \(B\) (obs2). Ifobs2is omitted, autocorrelation ofobs1is calculated by default.obs2 (
espressomd.observables.Observable, optional) – The observable \(B\) to be correlated with \(A\) (obs1).corr_operation (
str) – The operation that is performed on \(A(t)\) and \(B(t+\tau)\).delta_N (
int) – Number of timesteps between subsequent samples for the auto update mechanism.tau_max (
float) – This is the maximum value of \(\tau\) for which the correlation should be computed. Warning: Unless you are using the multiple tau correlator, choosingtau_maxof more than100 * dtwill result in a huge computational overhead. In a multiple tau correlator with reasonable parameters,tau_maxcan span the entire simulation without too much additional cpu time.tau_lin (
int) – The number of data-points for which the results are linearly spaced intau. This is a parameter of the multiple tau correlator. If you want to use it, make sure that you know how it works.tau_linmust be divisible by 2. By settingtau_linsuch thattau_max >= dt * delta_N * tau_lin, the multiple tau correlator is used, otherwise the trivial linear correlator is used. By settingtau_lin = 1, the value will be overridden bytau_lin = ceil(tau_max / (dt * delta_N)), which will result in either the multiple or linear tau correlator. In many cases,tau_lin=16is a good choice but this may strongly depend on the observables you are correlating. For more information, we recommend to read ref. [Ramírez et al., 2010] or to perform your own tests.compress1 (
str) – These functions are used to compress the data when going to the next level of the multiple tau correlator. This is done by producing one value out of two. The following compression functions are available:"discard2": (default value) discard the second value from the time series, use the first value as the result"discard1": discard the first value from the time series, use the second value as the result"linear": make a linear combination (average) of the two values
If only
compress1is specified, then the same compression function is used for both observables. If bothcompress1andcompress2are specified, thencompress1is used forobs1andcompress2forobs2.Both
discard1anddiscard2are safe for all observables but produce poor statistics in the tail. For some observables,"linear"compression can be used which makes an average of two neighboring values but produces systematic errors. Depending on the observable, the systematic error using the"linear"compression can be anything between harmless and disastrous. For more information, we recommend to read ref. [Ramírez et al., 2010] or to perform your own tests.compress2 (
str, optional) – Seecompress1.args (
floatof length 3) – Three floats which are passed as arguments to the correlation function. Currently it is only used by"fcs_acf", which will square these values in the core; if you later decide to update these weights withobs.args = [...], you’ll have to provide already squared values! Other correlation operations will ignore these values.
- class espressomd.accumulators.MeanVarianceCalculator(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperAccumulates results from observables.
- Parameters:
delta_N (
int) – Number of timesteps between subsequent samples for the auto update mechanism.
- update()¶
Update the accumulator (get the current values from the observable).
- mean()[source]¶
Returns the samples mean values of the respective observable with which the accumulator was initialized.
- class espressomd.accumulators.TimeSeries(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperRecords results from observables.
- Parameters:
delta_N (
int) – Number of timesteps between subsequent samples for the auto update mechanism.
- update()¶
Update the accumulator (get the current values from the observable).
- clear()¶
Clear the data
espressomd.analyze module¶
- class espressomd.analyze.Analysis(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper- linear_momentum()¶
Calculates the system’s linear momentum.
- Parameters:
include_particles (
bool, optional) – Whether to include the particles contribution to the linear momentum. True by default.include_lbfluid (
bool, optional) – whether to include the lattice-Boltzmann fluid contribution to the linear momentum. True by default.
- Returns:
The linear momentum of the system.
- Return type:
(3,) array_like of
float
- center_of_mass()¶
Calculate the center of mass of particles of a given type.
Note that virtual sites are not included, as they do not have a meaningful mass.
- Parameters:
p_type (
int) – Particletypefor which to calculate the center of mass.- Returns:
The center of mass of the particles.
- Return type:
(3,) array_like of
float
- nbhood()¶
Get all particles in a defined neighborhood.
- Parameters:
pos (array of
float) – Reference position for the neighborhood.r_catch (
float) – Radius of the region.
- Returns:
The neighbouring particle ids.
- Return type:
(N,) array_like of
int
- particle_neighbor_pids()¶
Get a list of all short-range neighbors for each particle.
- Returns:
A dictionary where each item is a pair of a particle id and its respective neighboring particle ids.
- Return type:
obj: dict
- calc_re()¶
Calculate the mean end-to-end distance of chains and its standard deviation, as well as mean square end-to-end distance of chains and its standard deviation.
This requires that a set of chains of equal length which start with the particle number
chain_startand are consecutively numbered, the last particle in that topology having id numberchain_start + number_of_chains * chain_length - 1.- Parameters:
chain_start (
int) – The id of the first monomer of the first chain.number_of_chains (
int) – Number of chains contained in the range.chain_length (
int) – The length of every chain.
- Returns:
Where [0] is the mean end-to-end distance of chains and [1] its standard deviation, [2] the mean square end-to-end distance and [3] its standard deviation.
- Return type:
(4,) array_like of
float
- calc_rg()¶
Calculate the mean radius of gyration of chains and its standard deviation, as well as the mean square radius of gyration of chains and its standard deviation.
This requires that a set of chains of equal length which start with the particle number
chain_startand are consecutively numbered, the last particle in that topology having id numberchain_start + number_of_chains * chain_length - 1.The radius of gyration is the radius of a sphere which would have the same moment of inertia as a polymer, and is defined as
\[R_{\mathrm G}^2 = \frac{1}{N} \sum\limits_{i=1}^{N} \left(\vec r_i - \vec r_{\mathrm{cm}}\right)^2\,,\]where \(\vec r_i\) are position vectors of individual particles constituting the polymer and \(\vec r_{\mathrm{cm}}\) is the position of its center of mass. The sum runs over all \(N\) particles comprising the polymer. For more information see any polymer science book, e.g. [Rubinstein and Colby, 2003].
- Parameters:
chain_start (
int) – The id of the first monomer of the first chain.number_of_chains (
int) – Number of chains contained in the range.chain_length (
int) – The length of every chain.
- Returns:
Where [0] is the mean radius of gyration of the chains and [1] its standard deviation, [2] the mean square radius of gyration and [3] its standard deviation.
- Return type:
(4,) array_like of
float
- calc_rh()¶
Calculate the mean hydrodynamic radius of chains and its standard deviation.
This requires that a set of chains of equal length which start with the particle number
chain_startand are consecutively numbered, the last particle in that topology having id numberchain_start + number_of_chains * chain_length - 1.The following formula is used for the calculation:
\[\frac{1}{R_{\mathrm H}} = \frac{2}{N(N-1)} \sum\limits_{i=1}^{N} \sum\limits_{j<i}^{N} \frac{1}{|\vec r_i - \vec r_j|}\,,\]This formula is only valid under certain assumptions. For more information, see chapter 4 and equation 4.102 in [Doi and Edwards, 1986]. Note that the hydrodynamic radius is sometimes defined in a similar fashion but with a denominator of \(N^2\) instead of \(N(N-1)\) in the prefactor. Both versions are equivalent in the \(N\rightarrow \infty\) limit but give numerically different values for finite polymers.
- Parameters:
chain_start (
int) – The id of the first monomer of the first chainnumber_of_chains (
int) – Number of chains contained in the range.chain_length (
int) – The length of every chain.
- Returns:
Where [0] is the mean hydrodynamic radius of the chains and [1] its standard deviation.
- Return type:
(2,) array_like of
float
- angular_momentum()¶
Calculate the system’s angular momentum with respect to the origin.
Note that virtual sites are not included, as they do not have a meaningful mass.
- Parameters:
p_type (
int) – Particletypefor which to calculate the center of mass, or-1for all particles.- Returns:
The center of mass of the system.
- Return type:
(3,) array_like of
float
- structure_factor()¶
Calculate the structure factor for given types. Returns the spherically averaged structure factor of particles specified in
sf_types. The structure factor is calculated for all possible wave vectors q up tosf_order. Do not choose parametersf_ordertoo large because the number of calculations grows assf_orderto the third power.- Parameters:
sf_types (list of
int) – Particletypewhich should be considered.sf_order (
int) – Specifies the maximum wavevector.
- Returns:
Where [0] contains the wave vectors q and [1] contains the structure factors S(q)
- Return type:
ndarray
- distribution()¶
Calculates the distance distribution of particles (probability of finding a particle of type A at a certain distance around a particle of type B, disregarding the fact that a spherical shell of a larger radius covers a larger volume). The distance is defined as the minimal distance between a particle of group
type_list_ato any of the grouptype_list_b. Returns two arrays, the bins and the (normalized) distribution.- Parameters:
type_list_a (list of
int) – List of particletype, only consider distances from these types.type_list_b (list of
int) – List of particletype, only consider distances to these types.r_min (
float) – Minimum distance. Default is 0.r_max (
float) – Maximum distance. By default, it is half the box size. A value larger than half the box size is allowed for systems with open boundary conditions.r_bins (
int) – Number of bins. Default is 100.log_flag (
bool) – When set toFalse, the bins are linearly equidistant; when set toTrue, the bins are logarithmically equidistant.int_flag (
bool) – When set toTrue, the result is an integrated distribution.
- Returns:
Where [0] contains the midpoints of the bins, and [1] contains the values of the rdf.
- Return type:
ndarray
- energy()[source]¶
Calculate the system energy in parallel.
- Returns:
A dictionary with the following keys:
"total": total energy"kinetic": linear and rotational kinetic energy"bonded": total bonded energy"bonded", <bond_id>: bonded energy from the bond identified bybond_id"non_bonded": total non-bonded energy"non_bonded", <type_i>, <type_j>: non-bonded energy from short-range interactions betweentype_iandtype_j"non_bonded_intra", <type_i>, <type_j>: non-bonded energy from short-range interactions betweentype_iandtype_jwith the samemol_id"non_bonded_inter", <type_i>, <type_j>: non-bonded energy from short-range interactions betweentype_iandtype_jwith differentmol_id"coulomb": Coulomb energy, how it is calculated depends on the method"coulomb", <i>: Coulomb energy from particle pairs (i=0), electrostatics solvers (i=1)"dipolar": dipolar energy"dipolar", <i>: dipolar energy from particle pairs and magnetic field constraints (i=0), magnetostatics solvers (i=1)"external_fields": external fields contribution
- Return type:
dict
Examples
>>> energy = system.analysis.energy() >>> print(energy["total"]) >>> print(energy["kinetic"]) >>> print(energy["bonded"]) >>> print(energy["non_bonded"]) >>> print(energy["external_fields"])
- gyration_tensor(p_type=None)[source]¶
Analyze the gyration tensor of particles of a given type.
- Parameters:
p_type (list of
int) – A particletype, or list of particle types to be considered.- Returns:
A dictionary with the following keys:
"Rg^2": squared radius of gyration"shape": three shape descriptors (asphericity, acylindricity, and relative shape anisotropy)"eva0": eigenvalue 0 of the gyration tensor and its corresponding eigenvector."eva1": eigenvalue 1 of the gyration tensor and its corresponding eigenvector."eva2": eigenvalue 2 of the gyration tensor and its corresponding eigenvector.
The eigenvalues are sorted in descending order.
- Return type:
dict
- min_dist(p1='default', p2='default')[source]¶
Minimal distance between two sets of particle types.
- Parameters:
p1, p2 (lists of
int) – Particletypein both sets. If both are set to'default', the minimum distance of all pairs is returned.- Returns:
Minimal distance.
- Return type:
float
- moment_of_inertia_matrix(p_type=None)[source]¶
Returns the 3x3 moment of inertia matrix for particles of a given type.
- Parameters:
p_type (
int) – A particletype.- Returns:
Moment of inertia matrix.
- Return type:
(3,3) array_like of
float
- particle_energy(particle)[source]¶
Calculate the non-bonded energy of a single given particle.
- Parameters:
particle (
ParticleHandle)- Returns:
Non-bonded energy of that particle
- Return type:
obj: float
- pressure()[source]¶
Calculate the instantaneous scalar pressure in parallel. This is only sensible in an isotropic system which is homogeneous (on average)! Do not use this in an anisotropic or inhomogeneous system. In order to obtain the pressure, the ensemble average needs to be calculated.
- Returns:
A dictionary with the following keys:
"total": total pressure"kinetic": kinetic pressure"bonded": total bonded pressure"bonded", <bond_id>: bonded pressure from the bond identified bybond_id"non_bonded": total non-bonded pressure"non_bonded", <type_i>, <type_j>: non-bonded pressure which arises from the interactions betweentype_iandtype_j"non_bonded_intra", <type_i>, <type_j>: non-bonded pressure from short-range forces betweentype_iandtype_jwith the samemol_id"non_bonded_inter", <type_i>, <type_j>: non-bonded pressure from short-range forces betweentype_iandtype_jwith differentmol_id"coulomb": Coulomb pressure, how it is calculated depends on the method. It is equivalent to 1/3 of the trace of the Coulomb pressure tensor. For how the pressure tensor is calculated, see Pressure Tensor. The averaged value in an isotropic NVT simulation is equivalent to the average of \(E^{\mathrm{coulomb}}/(3V)\), see [Brown and Neyertz, 1995]."coulomb", <i>: Coulomb pressure from particle pairs (i=0), electrostatics solvers (i=1)"dipolar": not implemented"virtual_sites": Pressure contribution from virtual sites"external_fields": external fields contribution
- Return type:
dict
- pressure_tensor()[source]¶
Calculate the instantaneous pressure tensor in parallel. This is sensible in an anisotropic system. Still it assumes that the system is homogeneous since the volume-averaged pressure tensor is used. Do not use this pressure tensor in an (on average) inhomogeneous system. If the system is (on average inhomogeneous) then use a local pressure tensor. In order to obtain the pressure tensor, the ensemble average needs to be calculated.
- Returns:
A dictionary with the following keys:
"total": total pressure tensor"kinetic": kinetic pressure tensor"bonded": total bonded pressure tensor"bonded", <bond_id>: bonded pressure tensor from the bond identified bybond_id"non_bonded": total non-bonded pressure tensor"non_bonded", <type_i>, <type_j>: non-bonded pressure tensor from short-range forces betweentype_iandtype_j"non_bonded_intra", <type_i>, <type_j>: non-bonded pressure tensor from short-range forces betweentype_iandtype_jwith the samemol_id"non_bonded_inter", <type_i>, <type_j>: non-bonded pressure tensor from short-range forces betweentype_iandtype_jwith differentmol_id"coulomb": Maxwell pressure tensor, how it is calculated depends on the method"coulomb", <i>: Maxwell pressure tensor from particle pairs (i=0), electrostatics solvers (i=1)"dipolar": not implemented"virtual_sites": pressure tensor contribution from virtual sites"external_fields": external fields contribution
- Return type:
dict
- espressomd.analyze.autocorrelation(time_series)[source]¶
Calculate the unnormalized autocorrelation function \(R_{XX}\) of an observable \(X\) measured at time \(t\) with constant time step for lag times \(\tau\) such that:
\(\displaystyle R_{XX}(\tau) = \frac{1}{N - \tau} \sum_{t=0}^{N - \tau} X_{t} \cdot X_{t + \tau}\)
This is a scipy implementation of the algorithm described in [de Buyl, 2018] that uses FFT. This implementation:
doesn’t subtract the mean of the time series before calculating the ACF,
doesn’t normalize the ACF by the variance of the time series,
assumes the time series is aperiodic (i.e. uses zero-padding).
- Parameters:
time_series ((N,) or (N, M) array_like of
float) – Time series to correlate. For multi-dimensional data, M is the number of columns.- Returns:
The time series autocorrelation function.
- Return type:
(N,) array_like of
float
espressomd.bond_breakage module¶
- class espressomd.bond_breakage.BreakageSpec(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperSpecifications for bond breakage. See Deleting bonds when particles are pulled apart for more details.
- Parameters:
breakage_length (
float) – Maximal bond extension until the bond breaks.action_type (
str, {‘delete_bond’, ‘revert_bind_at_point_of_collision’, ‘none’}) – Action triggered when the bond reaches its maximal extension.
- class espressomd.bond_breakage.BreakageSpecs(**kwargs)[source]¶
Bases:
ScriptObjectMap
espressomd.cell_system module¶
- class espressomd.cell_system.CellSystem(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperThis class controls the particle decomposition.
- decomposition_type¶
Name of the currently active particle decomposition.
- Type:
str
- use_verlet_lists¶
Whether to use Verlet lists.
- Type:
bool
- skin¶
Verlet list skin.
- Type:
float
- node_grid¶
MPI repartition for the regular decomposition cell system.
- Type:
(3,) array_like of
int
- max_cut_bonded¶
Maximal range from bonded interactions.
- Type:
float
- max_cut_nonbonded¶
Maximal range from non-bonded interactions.
- Type:
float
- interaction_range¶
Maximal interaction range from all interactions, or -1 when no interactions are active (or their cutoff has no impact when only 1 MPI rank is used).
- Type:
float
- resort()¶
Resort the particles in the cell system.
- Parameters:
global_flag (
bool, optional) – If true (default), a global resorting is done, otherwise particles are only exchanged between neighboring nodes.- Returns:
The number of particles per node.
- Return type:
(N,) array_like of
int
- tune_skin()¶
Tune the skin by measuring the integration time and bisecting over the given range of skins. The best skin is set in the simulation core.
- Parameters:
min_skin (
float) – Minimum skin to test.max_skin (
float) – Maximum skin.tol (
float) – Accuracy in skin to tune to.int_steps (
int) – Integration steps to time.adjust_max_skin (
bool, optional) – IfTrue, the value ofmax_skinis reduced to the maximum permissible skin (in case the passed value is too large). Defaults toFalse.
- Returns:
The
skin- Return type:
float
- get_state()¶
Get the current state of the cell system.
- Returns:
The cell system state.
- Return type:
dict
- get_neighbors(particle, distance)[source]¶
Get neighbors of a given particle up to a certain distance.
The choice of cell systems has an impact on how far the algorithm can detect particle pairs:
N-square: no restriction on the search distance, no double counting if search distance is larger than the box size
regular decomposition: the search distance is bounded by half the local cell geometry
hybrid decomposition: not supported
- Parameters:
particle (
ParticleHandle)distance (
float) – Pairs of particles closer thandistanceare found.
- Returns:
The list of neighbor particles surrounding the particle
- Return type:
(N,) array_like of
int
- get_pairs(distance, types='all')[source]¶
Get pairs of particles closer than threshold value.
- Parameters:
distance (
float) – Pairs of particles closer thandistanceare found.types (list of
intor'all', optional) – Restrict the pair search to the specified types. Defaults to'all', in which case all particles are considered.
- Returns:
The particle pairs identified by their index
- Return type:
list of tuples of
int- Raises:
Exception – If the pair search distance is greater than the cell size
- set_hybrid_decomposition(**kwargs)[source]¶
Activate the hybrid domain decomposition.
- Parameters:
cutoff_regular (
float) – Maximum cutoff to consider in regular decomposition. Should be as low as the system permits.n_square_types (list of
int, optional) – Particles types that should be handled in the N-square cell system. Defaults to an empty list.use_verlet_lists (
bool, optional) – Activates or deactivates the usage of Verlet lists. Defaults toTrue.
espressomd.checkpointing module¶
- class espressomd.checkpointing.Checkpoint(checkpoint_id=None, checkpoint_path='.')[source]¶
Bases:
objectCheckpoint handling (reading and writing).
- Parameters:
checkpoint_id (
str) – A string identifying a specific checkpoint.checkpoint_path (
str, optional) – Path for reading and writing the checkpoint. If not given, the current working directory is used.
- get_last_checkpoint_index()[source]¶
Returns the last index of the given checkpoint id. Will raise exception if no checkpoints are found.
- get_registered_objects()[source]¶
Returns a list of all object names that are registered for checkpointing.
- has_checkpoints()[source]¶
Check for checkpoints.
- Returns:
Trueif any checkpoints exist that matchcheckpoint_idandcheckpoint_pathotherwiseFalse.- Return type:
bool
- load(checkpoint_index=None)[source]¶
Loads the python objects using (c)Pickle and sets them in the calling module.
- Parameters:
checkpoint_index (
int, optional) – If not given, the lastcheckpoint_indexwill be used.
- read_signals()[source]¶
Reads all registered signals from the signal file and returns a list of integers.
- register(*args)[source]¶
Register python objects for checkpointing.
- Parameters:
args (list of
str) – Names of python objects to be registered for checkpointing.
- register_signal(signum=None)[source]¶
Register a signal that will trigger the signal handler.
- Parameters:
signum (
int) – Signal to be registered.
espressomd.cluster_analysis module¶
- class espressomd.cluster_analysis.Cluster(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperClass representing a cluster of particles.
- particle_ids()¶
Returns list of particle ids in the cluster
- size()¶
Returns the number of particles in the cluster
- center_of_mass()¶
Center of mass of the cluster (folded coordinates)
- longest_distance()¶
Longest distance between any combination of two particles in the cluster
- fractal_dimension(dr=None)¶
Estimates the cluster’s fractal dimension by fitting the number of particles \(n\) in spheres of growing radius around the center of mass to \(c*r_g^d\), where \(r_g\) is the radius of gyration of the particles within the sphere, and \(d\) is the fractal dimension.
Note
Requires
GSLexternal feature, enabled with-D ESPRESSO_BUILD_WITH_GSL=ON.- Parameters:
dr (
float) – Minimum increment for the radius of the spheres.- Returns:
Fractal dimension and mean square residual.
- Return type:
tuple
- class espressomd.cluster_analysis.ClusterStructure(*args, **kwargs)[source]¶
Bases:
ScriptInterfaceHelperCluster structure of a simulation system, and access to cluster analysis
- Parameters:
pair_criterion (
espressomd.pair_criteria._PairCriterion) – Criterion to decide whether two particles are neighbors.
- cid_for_particle(p)[source]¶
Returns cluster id for the particle.
- Parameters:
p (
espressomd.particle_data.ParticleHandleorintcontaining the particle id) – Particle.
- property clusters¶
Gives access to the clusters in the cluster structure via an instance of
Clusters.
- class espressomd.cluster_analysis.Clusters(cluster_structure)[source]¶
Bases:
objectAccess to the clusters in the cluster structure.
Access is as follows:
number of clusters:
len(clusters)access a cluster via its id:
clusters[id]iterate over clusters (yields
(id, cluster)tuples)
Example:
>>> for cluster_id, cluster in clusters: ... print(f"{cluster_id=} CoM={cluster.center_of_mass()}") cluster_id=1 CoM=[1.71834061 1.54988961 1.54734631]
espressomd.code_features module¶
- espressomd.code_features.assert_features(*args)[source]¶
Raise an exception when a list of features is not a subset of the compiled-in features.
espressomd.code_info module¶
- espressomd.code_info.all_features()[source]¶
Get the list of all features that can be activated in ESPResSo.
- espressomd.code_info.build_type()[source]¶
Get the CMake build type of this build of ESPResSo. Can be e.g. Debug, Release, RelWithAssert, RelWithDebInfo, Coverage, etc.
espressomd.collision_detection module¶
- class espressomd.collision_detection.CollisionDetection(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperInterface to the collision detection / dynamic binding.
See Creating bonds when particles collide for detailed instructions.
This class should not be instantiated by the user. Instead, use the
collision_detectionattribute of the system class to access the collision detection.Use method
set_params()to change the parameters of the collision detection.- set_params(**kwargs)[source]¶
Set the parameters for the collision detection
See Creating bonds when particles collide for detailed instructions.
- Parameters:
mode (
str, {“off”, “bind_centers”, “bind_at_point_of_collision”, “bind_three_particles”, “glue_to_surface”}) – Collision detection modedistance (
float) – Distance below which a pair of particles is considered in the collision detectionbond_centers (
espressomd.interactions.BondedInteraction) – Bond to add between the colliding particlesbond_vs (
espressomd.interactions.BondedInteraction) – Bond to add between virtual sites (for modes using virtual sites)part_type_vs (
int) – Particle type of the virtual sites being created on collision (virtual sites based modes)part_type_to_be_glued (
int) – particle type for"glue_to_surface"mode. See user guide.part_type_to_attach_vs_to (
int) – particle type for"glue_to_surface"mode. See user guide.part_type_after_glueing (
int) – particle type for"glue_to_surface"mode. See user guide.distance_glued_particle_to_vs (
float) – Distance for"glue_to_surface"mode. See user guide.bond_three_particles (
espressomd.interactions.BondedInteraction) – First angular bond for the"bind_three_particles"mode. See user guidethree_particle_binding_angle_resolution (
int) – Resolution for the angular bonds (mode"bind_three_particles"). Resolution+1 bonds are needed to accommodate the case of 180 degrees angles
espressomd.comfixed module¶
- class espressomd.comfixed.ComFixed(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperFix the center of mass of specific types.
Subtracts mass-weighted fraction of the total force action on all particles of the type from the particles after each force calculation. This keeps the center of mass of the type fixed iff the total momentum of the type is zero.
- Parameters:
types (array_like) – List of types for which the center of mass should be fixed.
espressomd.constraints module¶
- class espressomd.constraints.Constraint(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperBase class for constraints. A constraint provides a force and an energy contribution for a single particle.
- class espressomd.constraints.Constraints(**kwargs)[source]¶
Bases:
ScriptObjectListList of active constraints. Add a
espressomd.constraints.Constraintto make it active in the system, or remove it to make it inactive.- add(*args, **kwargs)[source]¶
Add a constraint to the list.
- Parameters:
constraint (
espressomd.constraints.Constraint) – Either a constraint object…**kwargs – … or parameters to construct an
espressomd.constraints.ShapeBasedConstraint
- Returns:
constraint – The added constraint
- Return type:
- remove(constraint)[source]¶
Remove a constraint from the list.
- Parameters:
constraint (
espressomd.constraints.Constraint)
- class espressomd.constraints.ElectricPlaneWave(phi=0, **kwargs)[source]¶
Bases:
ConstraintElectric field of the form
\(\vec{E} = \vec{E_0} \cdot \sin(\vec{k} \cdot \vec{x} + \omega \cdot t + \phi)\)
The resulting force on the particles are then
\(\vec{F} = q \cdot \vec{E}\)
where \(q\) and \(\vec{x}\) are the particle charge and position in folded coordinates. This can be used to generate a homogeneous AC field by setting \(\vec{k}\) to the null vector. For periodic systems, \(\vec{k}\) must be an integer multiple of \(2\pi \vec{L}^{-1}\) with \(\vec{L}\) the box length.
- Parameters:
E0 (array_like of
float) – Amplitude of the electric field.k (array_like of
float) – Wave vector of the waveomega (
float) – Frequency of the wavephi (
float, optional) – Phase
- property E0¶
- property k¶
- property omega¶
- property phi¶
- class espressomd.constraints.ElectricPotential(**kwargs)[source]¶
Bases:
_InterpolatedElectric potential interpolated from provided data. The electric field \(\vec{E}\) is calculated numerically from the potential, and the resulting force on the particles are
\(\vec{F} = q \cdot \vec{E}\)
where \(q\) is the charge of the particle.
- Parameters:
field ((M, N, O, 1) array_like of
float) – Potential on a grid of size (M, N, O)grid_spacing ((3,) array_like of
float) – Spacing of the grid points.
- class espressomd.constraints.FlowField(**kwargs)[source]¶
Bases:
_InterpolatedViscous coupling to a flow field that is interpolated from tabulated data like
\(\vec{F} = -\gamma \cdot \left( \vec{u}(\vec{x}) - \vec{v} \right)\)
where \(\vec{v}\) and \(\vec{x}\) are the particle velocity and position in folded coordinates, and \(\vec{u}(\vec{x})\) is a 3D flow field on a grid.
- Parameters:
field ((M, N, O, 3) array_like of
float) – Field velocity on a grid of size (M, N, O)grid_spacing ((3,) array_like of
float) – Spacing of the grid points.gamma (
float) – Coupling constant
- class espressomd.constraints.ForceField(**kwargs)[source]¶
Bases:
_InterpolatedA generic tabulated force field that applies a per-particle scaling factor.
- Parameters:
field ((M, N, O, 3) array_like of
float) – Forcefield amplitude on a grid of size (M, N, O).grid_spacing ((3,) array_like of
float) – Spacing of the grid points.default_scale (
float) – Scaling factor for particles that have no individual scaling factor.particle_scales (
dict) – A dictionary mapping particle ids to scaling factors. For these particles, the interaction is scaled with their individual scaling factor. Other particles are scaled with the default scaling factor.
- class espressomd.constraints.Gravity(**kwargs)[source]¶
Bases:
ConstraintGravity force
\(\vec{F} = m \cdot \vec{g}\)
- Parameters:
g ((3,) array_like of
float) – The gravitational constant.
- property g¶
- class espressomd.constraints.HomogeneousFlowField(**kwargs)[source]¶
Bases:
ConstraintViscous coupling to a flow field that is constant in space with the force
\(\vec{F} = -\gamma \cdot (\vec{u} - \vec{v})\)
where \(\vec{v}\) is the velocity of the particle and \(\vec{u}\) is the constant flow field.
- gamma¶
Coupling constant
- Type:
float
- property u¶
Field velocity ((3,) array_like of
float).
- class espressomd.constraints.HomogeneousMagneticField(**kwargs)[source]¶
Bases:
ConstraintHomogeneous magnetic field \(\vec{H}\). The resulting force \(\vec{F}\), torque \(\vec{\tau}\) and energy U on the particles are then
\(\vec{F} = \vec{0}\)
\(\vec{\tau} = \vec{\mu} \times \vec{H}\)
\(U = -\vec{\mu} \cdot \vec{H}\)
where \(\vec{\mu}\) is the particle dipole moment.
- H¶
Magnetic field vector. Describes both field direction and strength of the magnetic field (via length of the vector).
- Type:
(3,) array_like of
float
- class espressomd.constraints.LinearElectricPotential(phi0=0, **kwargs)[source]¶
Bases:
ConstraintElectric potential of the form
\(\phi = -\vec{E} \cdot \vec{x} + \phi_0\),
resulting in the electric field \(\vec{E}\) everywhere. The resulting force on the particles are then
\(\vec{F} = q \cdot \vec{E}\)
where \(q\) and \(\vec{x}\) are the particle charge and position in folded coordinates. This can be used to model a plate capacitor.
- Parameters:
E (array_like of
float) – The electric field.phi0 (
float) – The potential at the origin
- property E¶
- property phi0¶
- class espressomd.constraints.PotentialField(**kwargs)[source]¶
Bases:
_InterpolatedA generic tabulated force field that applies a per-particle scaling factor. The forces are calculated numerically from the data by finite differences. The potential is interpolated from the provided data.
- Parameters:
field ((M, N, O, 1) array_like of
float) – Potential on a grid of size (M, N, O).grid_spacing ((3,) array_like of
float) – Spacing of the grid points.default_scale (
float) – Scaling factor for particles that have no individual scaling factor.particle_scales (
dict) – A dictionary mapping particle ids to scaling factors. For these particles, the interaction is scaled with their individual scaling factor. Other particles are scaled with the default scaling factor.
- class espressomd.constraints.ShapeBasedConstraint(**kwargs)[source]¶
Bases:
Constraint- only_positive¶
Act only in the direction of positive normal, only useful if penetrable is
True.- Type:
bool
- particle_type¶
Interaction type of the constraint.
- Type:
int
- particle_velocity¶
Interaction velocity of the boundary
- Type:
array_like of
float
- penetrable¶
Whether particles are allowed to penetrate the constraint.
- Type:
bool
- shape¶
One of the shapes from
espressomd.shapes- Type:
See also
espressomd.shapesshape module that defines mathematical surfaces
Examples
>>> import espressomd >>> import espressomd.shapes >>> system = espressomd.System(box_l=3 * [10.]) >>> >>> # create first a shape-object to define the constraint surface >>> spherical_cavity = espressomd.shapes.Sphere(center=system.box_l / 2, radius=2.0, direction=-1.0) >>> >>> # now create an un-penetrable shape-based constraint of type 0 >>> spherical_constraint = system.constraints.add(particle_type=0, penetrable=False, shape=spherical_cavity) >>> >>> # place a trapped particle inside this sphere >>> system.part.add(pos=0.51 * system.box_l, type=1)
- min_dist()[source]¶
Calculates the minimum distance to all interacting particles.
- Returns:
The minimum distance
- Return type:
float
- total_force()[source]¶
Get total force acting on this constraint.
Examples
>>> import espressomd >>> import espressomd.shapes >>> system = espressomd.System(box_l=[50., 50., 50.]) >>> system.time_step = 0.01 >>> system.thermostat.set_langevin(kT=0.0, gamma=1.0) >>> system.cell_system.set_n_square(use_verlet_lists=False) >>> system.non_bonded_inter[0, 0].lennard_jones.set_params( ... epsilon=1, sigma=1, cutoff=2**(1. / 6), shift="auto") >>> >>> floor = system.constraints.add( ... shape=espressomd.shapes.Wall(normal=[0, 0, 1], dist=0.0), ... particle_type=0, penetrable=False, only_positive=False) >>> >>> p = system.part.add(pos=[0,0,1.5], type=0, ext_force=[0, 0, -.1]) >>> # print the particle position as it falls >>> # and print the force it applies on the floor >>> for t in range(10): ... system.integrator.run(100) ... print(p.pos, floor.total_force())
espressomd.cuda_init module¶
- class espressomd.cuda_init.CudaInitHandle(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper- device¶
Device id to use.
- Type:
int
- list_devices()¶
List devices.
- Returns:
Available CUDA devices sorted by device id.
- Return type:
dict
espressomd.drude_helpers module¶
- class espressomd.drude_helpers.DrudeHelpers[source]¶
Bases:
objectProvides helper functions to assist in the creation of Drude particles.
- add_all_thole(system, verbose=False)[source]¶
Calls
add_thole_pair_damping()for all necessary combinations to create the interactions.- Parameters:
system (
espressomd.system.System)verbose (
bool) – Turns on verbosity.
- add_drude_particle_to_core(system, harmonic_bond, thermalized_bond, p_core, type_drude, alpha, mass_drude, coulomb_prefactor, thole_damping=2.6, verbose=False)[source]¶
Adds a Drude particle with specified type and mass to the system and returns its espressomd.particle_data.ParticleHandle. Checks if different Drude particles have different types. Collects types/charges/polarizations/Thole factors for intramolecular core-Drude short-range exclusion and Thole interaction.
- Parameters:
system (
espressomd.system.System)harmonic_bond (
espressomd.interactions.HarmonicBond) – Add this harmonic bond to between Drude particle and corethermalized_bond (
espressomd.interactions.ThermalizedBond) – Add this thermalized_bond to between Drude particle and corep_core (
espressomd.particle_data.ParticleHandle) – The existing core particletype_drude (
int) – The type of the newly created Drude particlealpha (
float) – The polarizability in units of inverse volume. Related to the charge of the Drude particle.mass_drude (
float) – The mass of the newly created Drude particlecoulomb_prefactor (
float) – Required to calculate the charge of the Drude particle.thole_damping (
float) – Thole damping factor of the Drude pair. Comes to effect ifadd_all_thole()method is used.verbose (
bool) – Turns on verbosity.
- Returns:
The created Drude Particle.
- Return type:
- add_intramol_exclusion_bonds(drude_parts, core_parts, verbose=False)[source]¶
Applies electrostatic short-range exclusion bonds for the given ids. Has to be applied for all molecules.
- Parameters:
system (
espressomd.system.System)drude_parts – List of Drude particles within a molecule.
core_parts – List of core particles within a molecule.
verbose (
bool) – Turns on verbosity.
- add_thole_pair_damping(system, t1, t2, verbose=False)[source]¶
Calculates mixed Thole factors depending on Thole damping and polarization. Adds non-bonded Thole interactions to the system.
- Parameters:
system (
espressomd.system.System)t1 (
int) – Type 1t2 (
int) – Type 2verbose (
bool) – Turns on verbosity.
- setup_and_add_drude_exclusion_bonds(system, verbose=False)[source]¶
Creates electrostatic short-range exclusion bonds for global exclusion between Drude particles and core charges and adds the bonds to the cores. Has to be called once after all Drude particles have been created.
- Parameters:
system (
espressomd.system.System)verbose (
bool) – Turns on verbosity.
- setup_intramol_exclusion_bonds(system, mol_drude_types, mol_core_types, mol_core_partial_charges, verbose=False)[source]¶
Creates electrostatic short-range exclusion bonds for intramolecular exclusion between Drude particles and partial charges of the cores. Has to be called once after all Drude particles have been created.
- Parameters:
system (
espressomd.system.System)mol_drude_types – List of types of Drude particles within the molecule
mol_core_types – List of types of core particles within the molecule
mol_core_partial_charges – List of partial charges of core particles within the molecule
verbose (
bool) – Turns on verbosity.
espressomd.electrokinetics module¶
- class espressomd.electrokinetics.DensityBoundary(density)[source]¶
Bases:
objectHold density information for the density boundary condition at a single node.
- class espressomd.electrokinetics.EKBulkReaction(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperReaction type that is applied everywhere in the domain.
- Parameters:
lattice (
espressomd.electrokinetics.LatticeWalberla) – Lattice object.tau (
float) – EK time step, must be an integer multiple of the MD time step.coefficient (
float) – Reaction rate constant of the reaction.reactants (array_like of
EKReactant) – Reactants that participate this reaction.
- class espressomd.electrokinetics.EKContainer(*args, **kwargs)[source]¶
Bases:
ScriptObjectListContainer object holding the
EKSpecies.- Parameters:
- clear()¶
Removes all species.
- add(ekspecies)[source]¶
Add an
EKSpeciesto the container.- Parameters:
ekspecies (
EKSpecies) – Species to be added.
- property reactions¶
rtype: Reactions-container of the current reactions (
EKReactions).
- class espressomd.electrokinetics.EKFFT(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperA FFT-based Poisson solver. Intrinsically assumes periodic boundary conditions.
- Parameters:
lattice (
espressomd.lb.LatticeWalberla) – Lattice object.permittivity (
float) – permittivity of the fluid \(\epsilon_0 \epsilon_{\mathrm{r}}\).single_precision (
bool, optional) – Use single-precision floating-point arithmetic.
- class espressomd.electrokinetics.EKIndexedReaction(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperReaction type that is applied only on specific cells in the domain. Can be used to model surface-reactions.
- Parameters:
lattice (
espressomd.electrokinetics.LatticeWalberla) – Lattice object.tau (
float) – EK time step, must be an integer multiple of the MD time step.coefficient (
float) – Reaction rate constant of the reaction.reactants (array_like of
EKReactant) – Reactants that participate this reaction.
- class espressomd.electrokinetics.EKNone(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperThe default Poisson solver. Imposes a null electrostatic potential everywhere.
- Parameters:
lattice (
espressomd.lb.LatticeWalberla) – Lattice object.single_precision (
bool, optional) – Use single-precision floating-point arithmetic.
- class espressomd.electrokinetics.EKReactant(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperReactant-object which specifies the contribution of a species to a reaction.
- Parameters:
ekspecies (
EKSpecies) – EK species to reactstoech_coeff (
float) – Stoechiometric coefficient of this species in the reaction. Products have positive coefficients whereas educts have negative ones.order (
float) – Partial-order of this species in the reaction.
- class espressomd.electrokinetics.EKReactions(**kwargs)[source]¶
Bases:
ScriptObjectListContainer object holding all EK-reactions that are considered.
- clear()¶
Remove all reactions.
- add(reaction)[source]¶
Add a reaction to the container.
- Parameters:
reaction (
EKBulkReactionorEKIndexedReaction) – Reaction to be added.
- remove(reaction)[source]¶
Remove a reaction from the container.
- Parameters:
reaction (
EKBulkReactionorEKIndexedReaction) – Reaction to be removed.
- class espressomd.electrokinetics.EKSpecies(*args, **kwargs)[source]¶
Bases:
ScriptInterfaceHelper,LatticeModelThe advection-diffusion-reaction method for chemical species using waLBerla.
- Parameters:
lattice (
espressomd.electrokinetics.LatticeWalberla) – Lattice object.tau (
float) – EK time step, must be an integer multiple of the MD time step.density (
float) – Species density.diffusion (
float) – Species diffusion coefficient.valency (
float) – Species valency.advection (
bool) – Whether to enable advection.friction_coupling (
bool) – Whether to enable friction coupling.ext_efield ((3,) array_like of
float, optional) – External electrical field.kT (
float, optional) – Thermal energy of the simulated heat bath (for thermalized species). Set it to 0 for an unthermalized species.single_precision (
bool, optional) – Use single-precision floating-point arithmetic.
- clear_density_boundaries()¶
Remove density boundary conditions.
- clear_flux_boundaries()¶
Remove flux boundary conditions.
- clear_boundaries()¶
Remove all boundary conditions.
- save_checkpoint()¶
Write EK densities and boundary conditions to a file.
- Parameters:
path (
str) – Destination file path.binary (
bool) – Whether to write in binary or ASCII mode.
- load_checkpoint()¶
Load EK densities and boundary conditions from a file.
- Parameters:
path (
str) – File path to read from.binary (
bool) – Whether to read in binary or ASCII mode.
- add_vtk_writer()¶
Attach a VTK writer.
- Parameters:
vtk (
espressomd.electrokinetics.VTKOutput) – VTK writer.
- remove_vtk_writer()¶
Detach a VTK writer.
- Parameters:
vtk (
espressomd.electrokinetics.VTKOutput) – VTK writer.
- clear_vtk_writers()¶
Detach all VTK writers.
- add_boundary_from_shape(shape, value, boundary_type)[source]¶
Set boundary conditions from a shape.
- Parameters:
shape (
espressomd.shapes.Shape) – Shape to rasterize.value ((O,) or (L, M, N, O) array_like of
float, optional) – Boundary numerical value. If a single value of shape(O,)is given, it will be broadcast to all nodes inside the shape, otherwiseL, M, Nmust be equal to the EK grid dimensions.boundary_type (Union[
DensityBoundary,) –FluxBoundary] (optional) Type of the boundary condition.
- class espressomd.electrokinetics.EKSpeciesNode(*args, **kwargs)[source]¶
Bases:
ScriptInterfaceHelper- property density¶
- property density_boundary¶
returns: *
DensityBoundary– If the node is a boundary node *None– If the node is not a boundary node
- property flux_boundary¶
returns: *
FluxBoundary– If the node is a boundary node *None– If the node is not a boundary node
- property index¶
- property is_boundary¶
- class espressomd.electrokinetics.EKSpeciesSlice(*args, **kwargs)[source]¶
Bases:
ScriptInterfaceHelper- property density¶
- property density_boundary¶
returns: * (N, M, L) array_like of
DensityBoundary– If the nodes are boundary nodes * (N, M, L) array_like ofNone– If the nodes are not boundary nodes
- property flux_boundary¶
returns: * (N, M, L) array_like of
FluxBoundary– If the nodes are boundary nodes * (N, M, L) array_like of None` – If the nodes are not boundary nodes
- property is_boundary¶
- class espressomd.electrokinetics.FluxBoundary(flux)[source]¶
Bases:
objectHold flux information for the flux boundary condition at a single node.
- class espressomd.electrokinetics.VTKOutput(*args, **kwargs)[source]¶
Bases:
VTKOutputBaseCreate a VTK writer.
Files are written to
<base_folder>/<identifier>/<prefix>_*.vtu. Summary is written to<base_folder>/<identifier>.pvd.Manual VTK callbacks can be called at any time to take a snapshot of the current state of the EK species.
Automatic VTK callbacks can be disabled at any time and re-enabled later. Please note that the internal VTK counter is no longer incremented when an automatic callback is disabled, which means the number of EK steps between two frames will not always be an integer multiple of
delta_N.- Parameters:
identifier (
str) – Name of the VTK writer.observables (
list, {‘density’,}) – List of observables to write to the VTK files.delta_N (
int) – Write frequency. If this value is 0 (default), the object is a manual VTK callback that must be triggered manually. Otherwise, it is an automatic callback that is added to the time loop and writes everydelta_NEK steps.base_folder (
str(optional), default is ‘vtk_out’) – Path to the output VTK folder.prefix (
str(optional), default is ‘simulation_step’) – Prefix for VTK files.
espressomd.electrostatic_extensions module¶
- class espressomd.electrostatic_extensions.ElectrostaticExtensions(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper
- class espressomd.electrostatic_extensions.ICC(**kwargs)[source]¶
Bases:
ElectrostaticExtensionsInterface to the induced charge calculation scheme for dielectric interfaces. See Dielectric interfaces with the ICC\star algorithm for more details.
- Parameters:
n_icc (
int) – Total number of ICC Particles.first_id (
int, optional) – ID of the first ICC Particle.convergence (
float, optional) – Abort criteria of the iteration. It corresponds to the maximum relative change of any of the interface particle’s charge.relaxation (
float, optional) – SOR relaxation parameter.ext_field (
float, optional) – Homogeneous electric field added to the calculation of dielectric boundary forces.max_iterations (
int, optional) – Maximal number of iterations.eps_out (
float, optional) – Relative permittivity of the outer region (where the particles are).normals ((
n_icc, 3) array_likefloat) – Normal vectors pointing into the outer region.areas ((
n_icc, ) array_likefloat) – Areas of the discretized surface.sigmas ((
n_icc, ) array_likefloat, optional) – Additional surface charge density in the absence of any charge induction.epsilons ((
n_icc, ) array_likefloat) – Dielectric constant associated to the areas.
espressomd.electrostatics module¶
- class espressomd.electrostatics.Container(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper
- class espressomd.electrostatics.DH(**kwargs)[source]¶
Bases:
ElectrostaticInteractionElectrostatics solver based on the Debye-Hueckel framework. See Debye-Hückel potential for more details.
- Parameters:
prefactor (
float) – Electrostatics prefactor (see (1)).kappa (
float) – Inverse Debye screening length.r_cut (
float) – Cutoff radius for this interaction.
- class espressomd.electrostatics.ELC(**kwargs)[source]¶
Bases:
ElectrostaticInteractionElectrostatics solver for systems with two periodic dimensions. See Electrostatic Layer Correction (ELC) for more details.
- Parameters:
actor (
P3M, required) – Base P3M actor.gap_size (
float, required) – The gap size gives the height \(h\) of the empty region between the system box and the neighboring artificial images. ESPResSo checks that the gap is empty and will throw an error if it isn’t. Therefore you should really make sure that the gap region is empty (e.g. with wall constraints).maxPWerror (
float, required) – The maximal pairwise error sets the least upper bound (LUB) error of the force between any two charges without prefactors (see the papers). The algorithm tries to find parameters to meet this LUB requirements or will throw an error if there are none.delta_mid_top (
float, optional) – Dielectric contrast \(\Delta_t\) between the upper boundary and the simulation box. Value between -1 and +1 (inclusive).delta_mid_bottom (
float, optional) – Dielectric contrast \(\Delta_b\) between the lower boundary and the simulation box. Value between -1 and +1 (inclusive).const_pot (
bool, optional) – Activate a constant electric potential between the top and bottom of the simulation box.pot_diff (
float, optional) – Ifconst_potis enabled, this parameter controls the applied voltage between the boundaries of the simulation box in the z-direction (at \(z = 0\) and \(z = L_z - h\)).neutralize (
bool, optional) – By default, ELC just as P3M adds a homogeneous neutralizing background to the system in case of a net charge. However, unlike in three dimensions, this background adds a parabolic potential across the slab [Ballenegger et al., 2009]. Therefore, under normal circumstances, you will probably want to disable the neutralization for non-neutral systems. This corresponds then to a formal regularization of the forces and energies [Ballenegger et al., 2009]. Also, if you add neutralizing walls explicitly as constraints, you have to disable the neutralization. When using a dielectric contrast or full metallic walls (delta_mid_top != 0ordelta_mid_bot != 0orconst_pot=True),neutralizeis overwritten and switched off internally. Note that the special case of non-neutral systems with a non-metallic dielectric jump (e.g.delta_mid_topordelta_mid_botin]-1,1[) is not covered by the algorithm and will throw an error.far_cut (
float, optional) – Cutoff radius, use with care, intended for testing purposes. When setting the cutoff directly, the maximal pairwise error is ignored.
- class espressomd.electrostatics.ElectrostaticInteraction(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperCommon interface for electrostatics solvers.
- Parameters:
prefactor (
float) – Electrostatics prefactor \(\frac{1}{4\pi\varepsilon_0\varepsilon_r}\)
- class espressomd.electrostatics.MMM1D(**kwargs)[source]¶
Bases:
ElectrostaticInteractionElectrostatics solver for systems with one periodic direction. See MMM1D for more details.
- Parameters:
prefactor (
float) – Electrostatics prefactor (see (1)).maxWPerror (
float) – Maximal pairwise error.far_switch_radius (
float, optional) – Radius where near-field and far-field calculation are switched.verbose (
bool, optional) – IfFalse, disable log output during tuning.timings (
int, optional) – Number of force calculations during tuning.check_neutrality (
bool, optional) – Raise a warning if the system is not electrically neutral when set toTrue(default).
- class espressomd.electrostatics.MMM1DGPU(**kwargs)[source]¶
Bases:
ElectrostaticInteractionElectrostatics solver with GPU support for systems with one periodic direction. See MMM1D on GPU for more details.
- Parameters:
prefactor (
float) – Electrostatics prefactor (see (1)).maxWPerror (
float) – Maximal pairwise error.far_switch_radius (
float, optional) – Radius where near-field and far-field calculation are switchedbessel_cutoff (
int, optional)timings (
int, optional) – Number of force calculations during tuning.check_neutrality (
bool, optional) – Raise a warning if the system is not electrically neutral when set toTrue(default).
- class espressomd.electrostatics.P3M(**kwargs)[source]¶
Bases:
_P3MBaseP3M electrostatics solver.
Particle–Particle–Particle–Mesh (P3M) is a Fourier-based Ewald summation method to calculate potentials in N-body simulation. See Coulomb P3M for more details.
- Parameters:
prefactor (
float) – Electrostatics prefactor (see (1)).accuracy (
float) – P3M tunes its parameters to provide this target accuracy.alpha (
float, optional) – The Ewald parameter.cao (
float, optional) – The charge-assignment order, an integer between 1 and 7.epsilon (
floatorstr, optional) – A positive number for the dielectric constant of the surrounding medium. Use'metallic'to set the dielectric constant of the surrounding medium to infinity (default).mesh (
intor (3,) array_like ofint, optional) – The number of mesh points in x, y and z direction. Use a single value for cubic boxes.mesh_off ((3,) array_like of
float, optional) – Mesh offset.r_cut (
float, optional) – The real space cutoff.tune (
bool, optional) – Used to activate/deactivate the tuning method on activation. Defaults toTrue.timings (
int) – Number of force calculations during tuning.verbose (
bool, optional) – IfFalse, disable log output during tuning.check_neutrality (
bool, optional) – Raise a warning if the system is not electrically neutral when set toTrue(default).check_complex_residuals (
bool, optional) – Raise a warning if the backward Fourier transform has non-zero complex residuals when set toTrue(default).
- class espressomd.electrostatics.P3MGPU(**kwargs)[source]¶
Bases:
_P3MBaseP3M electrostatics solver with GPU support.
Particle–Particle–Particle–Mesh (P3M) is a Fourier-based Ewald summation method to calculate potentials in N-body simulation. See Coulomb P3M on GPU for more details.
- Parameters:
prefactor (
float) – Electrostatics prefactor (see (1)).accuracy (
float) – P3M tunes its parameters to provide this target accuracy.alpha (
float, optional) – The Ewald parameter.cao (
float, optional) – The charge-assignment order, an integer between 0 and 7.epsilon (
floatorstr, optional) – A positive number for the dielectric constant of the surrounding medium. Use'metallic'to set the dielectric constant of the surrounding medium to infinity (default).mesh (
intor (3,) array_like ofint, optional) – The number of mesh points in x, y and z direction. Use a single value for cubic boxes.mesh_off ((3,) array_like of
float, optional) – Mesh offset.r_cut (
float, optional) – The real space cutofftune (
bool, optional) – Used to activate/deactivate the tuning method on activation. Defaults toTrue.timings (
int) – Number of force calculations during tuning.verbose (
bool, optional) – IfFalse, disable log output during tuning.check_neutrality (
bool, optional) – Raise a warning if the system is not electrically neutral when set toTrue(default).check_complex_residuals (
bool, optional) – Raise a warning if the backward Fourier transform has non-zero complex residuals when set toTrue(default).
- class espressomd.electrostatics.ReactionField(**kwargs)[source]¶
Bases:
ElectrostaticInteractionElectrostatics solver based on the Reaction Field framework. See Reaction Field method for more details.
- Parameters:
prefactor (
float) – Electrostatics prefactor (see (1)).kappa (
float) – Inverse Debye screening length.epsilon1 (
float) – interior dielectric constantepsilon2 (
float) – exterior dielectric constantr_cut (
float) – Cutoff radius for this interaction.
- class espressomd.electrostatics.Scafacos(**kwargs)[source]¶
Bases:
ElectrostaticInteractionCalculate the Coulomb interaction using the ScaFaCoS library. See ScaFaCoS electrostatics for more details.
- Parameters:
prefactor (
float) – Coulomb prefactor as defined in (1).method_name (
str) – Name of the ScaFaCoS method to use.method_params (
dict) – Dictionary containing the method-specific parameters.
- get_available_methods()¶
List long-range methods available in the ScaFaCoS library.
- set_near_field_delegation()¶
Choose whether to delegate short-range calculation to ESPResSo (this is the default when the method supports it) or ScaFaCos.
- Parameters:
delegate (
bool) – Delegate to ESPResSo ifTrueand the method supports it.
- get_near_field_delegation()¶
Find whether the short-range calculation is delegated to ESPResSo (this is the default when the method supports it) or ScaFaCos.
- Returns:
delegate – Delegate to ESPResSo if
Trueand the method supports it,Falseif delegated to ScaFaCoS or the method doesn’t have a short-range kernel.- Return type:
bool
espressomd.galilei module¶
- class espressomd.galilei.GalileiTransform(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperCollective operations on particles.
- kill_particle_motion()¶
Stop the motion of all particles.
- Parameters:
rotation (
bool, optional) – Whether or not to stop the angular momentum too. Defaults to false.
- kill_particle_forces()¶
Set the forces on the particles to zero.
- Parameters:
torque (
bool, optional) – Whether or not to set the torques to zero too. Defaults to false.
- system_CMS()¶
Calculate the center of mass of the system. Assumes equal unit mass if the
MASSfeature is not compiled in.- Returns:
The center of mass position.
- Return type:
(3,) array_like of
float
- system_CMS_velocity()¶
Calculate the center of mass velocity of the system. Assumes equal unit mass if the
MASSfeature is not compiled in.- Returns:
The of the center of mass velocity vector.
- Return type:
(3,) array_like of
float
- galilei_transform()¶
Remove the center of mass velocity of the system. Assumes equal unit mass if the
MASSfeature is not compiled in. This is often used when switching from Langevin Dynamics to lattice-Boltzmann. This is due to the random nature of LD that yield a non-zero net system momentum at any given time.
espressomd.highlander module¶
espressomd.integrate module¶
- class espressomd.integrate.BrownianDynamics(**kwargs)[source]¶
Bases:
IntegratorBrownian Dynamics integrator.
- class espressomd.integrate.Integrator(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperIntegrator class.
- run(steps=1, recalc_forces=False, reuse_forces=False)[source]¶
Run the integrator.
- Parameters:
steps (
int) – Number of time steps to integrate.recalc_forces (
bool, optional) – Recalculate the forces regardless of whether they are reusable.reuse_forces (
bool, optional) – Reuse the forces from previous time step.
- class espressomd.integrate.IntegratorHandle(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperProvide access to the currently active integrator.
- set_isotropic_npt(**kwargs)[source]¶
Set the integration method to a modified velocity Verlet designed for simulations in the NpT ensemble (
VelocityVerletIsotropicNPT).
- set_nvt()[source]¶
Set the integration method to velocity Verlet, which is suitable for simulations in the NVT ensemble (
VelocityVerlet).
- set_steepest_descent(**kwargs)[source]¶
Set the integration method to steepest descent (
SteepestDescent).
- set_stokesian_dynamics(**kwargs)[source]¶
Set the integration method to Stokesian Dynamics (
StokesianDynamics).
- set_vv()[source]¶
Set the integration method to velocity Verlet, which is suitable for simulations in the NVT ensemble (
VelocityVerlet).
- class espressomd.integrate.SteepestDescent(**kwargs)[source]¶
Bases:
IntegratorSteepest descent algorithm for energy minimization.
Particles located at \(\vec{r}_i\) at integration step \(i\) and experiencing a potential \(\mathcal{H}(\vec{r}_i)\) are displaced according to the equation:
\(\vec{r}_{i+1} = \vec{r}_i - \gamma\nabla\mathcal{H}(\vec{r}_i)\)
- Parameters:
f_max (
float) – Convergence criterion. Minimization stops when the maximal force on particles in the system is lower than this threshold. Set this to 0 when running minimization in a loop that stops when a custom convergence criterion is met.gamma (
float) – Dampening constant.max_displacement (
float) – Maximal allowed displacement per step. Typical values for a LJ liquid are in the range of 0.1% to 10% of the particle sigma.
- class espressomd.integrate.StokesianDynamics(**kwargs)[source]¶
Bases:
IntegratorStokesian Dynamics integrator.
- Parameters:
viscosity (
float) – Bulk viscosity.radii (
dict) – Dictionary that maps particle types to radii.approximation_method (
str, optional, {‘ft’, ‘fts’}) – Chooses the method of the mobility approximation.'fts'is more accurate. Default is'fts'.self_mobility (
bool, optional) – Switches off or on the mobility terms for single particles. Default isTrue.pair_mobility (
bool, optional) – Switches off or on the hydrodynamic interactions between particles. Default isTrue.
- class espressomd.integrate.VelocityVerlet(**kwargs)[source]¶
Bases:
IntegratorVelocity Verlet integrator, suitable for simulations in the NVT ensemble.
- class espressomd.integrate.VelocityVerletIsotropicNPT(**kwargs)[source]¶
Bases:
IntegratorModified velocity Verlet integrator, suitable for simulations in the NpT ensemble with isotropic rescaling. Requires the NpT thermostat, activated with
espressomd.thermostat.Thermostat.set_npt().- Parameters:
ext_pressure (
float) – The external pressure.piston (
float) – The mass of the applied piston.direction ((3,) array_like of
bool, optional) – Select which dimensions are allowed to fluctuate by assigning them toTrue. Default is allTrue.cubic_box (
bool, optional) – IfTrue, a cubic box is assumed and the value ofdirectionwill be ignored when rescaling the box. This is required e.g. for electrostatics and magnetostatics. Default isFalse.
espressomd.interactions module¶
- class espressomd.interactions.AngleCosine(**kwargs)[source]¶
Bases:
BondedInteractionBond-angle-dependent cosine potential.
- Parameters:
phi0 (
float) – Equilibrium bond angle in radians.bend (
float) – Magnitude of the bond interaction.
- class espressomd.interactions.AngleCossquare(**kwargs)[source]¶
Bases:
BondedInteractionBond-angle-dependent cosine squared potential.
- Parameters:
phi0 (
float) – Equilibrium bond angle in radians.bend (
float) – Magnitude of the bond interaction.
- class espressomd.interactions.AngleHarmonic(**kwargs)[source]¶
Bases:
BondedInteractionBond-angle-dependent harmonic potential.
- Parameters:
phi0 (
float) – Equilibrium bond angle in radians.bend (
float) – Magnitude of the bond interaction.
- class espressomd.interactions.BMHTFInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionBMHTF interaction.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
a (
float) – Magnitude of exponential part of the interaction.b (
float) – Exponential factor of the interaction.c (
float) – Magnitude of the term decaying with the sixth power of r.d (
float) – Magnitude of the term decaying with the eighth power of r.sig (
float) – Shift in the exponent.cutoff (
float) – Cutoff distance of the interaction.
- class espressomd.interactions.BONDED_IA(value, names=<not given>, *values, module=None, qualname=None, type=None, start=1, boundary=None)[source]¶
Bases:
IntEnum- ANGLE_COSINE = 7¶
- ANGLE_COSSQUARE = 8¶
- ANGLE_HARMONIC = 6¶
- BONDED_COULOMB = 4¶
- BONDED_COULOMB_SR = 5¶
- DIHEDRAL = 9¶
- FENE = 1¶
- HARMONIC = 2¶
- IBM_TRIBEND = 17¶
- IBM_TRIEL = 15¶
- IBM_VOLUME_CONSERVATION = 16¶
- NONE = 0¶
- OIF_GLOBAL_FORCES = 18¶
- OIF_LOCAL_FORCES = 19¶
- QUARTIC = 3¶
- RIGID_BOND = 14¶
- TABULATED_ANGLE = 11¶
- TABULATED_DIHEDRAL = 12¶
- TABULATED_DISTANCE = 10¶
- THERMALIZED_DIST = 13¶
- VIRTUAL_BOND = 20¶
- class espressomd.interactions.BondedCoulomb(**kwargs)[source]¶
Bases:
BondedInteractionBonded Coulomb bond.
- Parameters:
prefactor (
float) – Coulomb prefactor of the bonded Coulomb interaction.
- class espressomd.interactions.BondedCoulombSRBond(**kwargs)[source]¶
Bases:
BondedInteractionBonded Coulomb short range bond. Calculates the short range part of Coulomb interactions.
- Parameters:
q1q2 (
float) – Charge factor of the involved particle pair. Note the particle charges are used to allow e.g. only partial subtraction of the involved charges.
- class espressomd.interactions.BondedInteraction(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperBase class for bonded interactions.
Either called with an interaction id, in which case the interaction will represent the bonded interaction as it is defined in ESPResSo core, or called with keyword arguments describing a new interaction.
- property params¶
- class espressomd.interactions.BondedInteractions(**kwargs)[source]¶
Bases:
ScriptObjectMapRepresents the bonded interactions list.
Individual interactions can be accessed using
BondedInteractions[i], whereiis the bond id.- remove()¶
Remove a bond from the list. This is a no-op if the bond does not exist.
- Parameters:
bond_id (
int)
- clear()¶
Remove all bonds.
- add(*args)[source]¶
Add a bond to the list.
- Parameters:
bond (
espressomd.interactions.BondedInteraction) – Either a bond object…
- class espressomd.interactions.BuckinghamInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionBuckingham interaction.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
a (
float) – Magnitude of the exponential part of the interaction.b (
float, optional) – Exponent of the exponential part of the interaction.c (
float) – Prefactor of term decaying with the sixth power of distance.d (
float) – Prefactor of term decaying with the fourth power of distance.discont (
float) – Distance below which the potential is linearly continued.cutoff (
float) – Cutoff distance of the interaction.shift (
float, optional) – Constant potential shift.
- class espressomd.interactions.DPDInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionDPD interaction.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
weight_function (
int, {0, 1}) – The distance dependence of the parallel part, either 0 (constant) or 1 (linear)gamma (
float) – Friction coefficient of the parallel partk (
float) – Exponent in the modified weight functionr_cut (
float) – Cutoff of the parallel parttrans_weight_function (
int, {0, 1}) – The distance dependence of the orthogonal part, either 0 (constant) or 1 (linear)trans_gamma (
float) – Friction coefficient of the orthogonal parttrans_r_cut (
float) – Cutoff of the orthogonal part
- class espressomd.interactions.Dihedral(**kwargs)[source]¶
Bases:
BondedInteractionDihedral potential with phase shift.
- Parameters:
mult (
int) – Multiplicity of the potential (number of minima).bend (
float) – Bending constant.phase (
float) – Angle of the first local minimum in radians.
- class espressomd.interactions.FeneBond(**kwargs)[source]¶
Bases:
BondedInteractionFENE bond.
- Parameters:
k (
float) – Magnitude of the bond interaction.d_r_max (
float) – Maximum stretch and compression length of the bond.r_0 (
float, optional) – Equilibrium bond length.
- class espressomd.interactions.GaussianInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionGaussian interaction.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
eps (
float) – Overlap energy epsilon.sig (
float) – Variance sigma of the Gaussian interaction.cutoff (
float) – Cutoff distance of the interaction.
- class espressomd.interactions.GayBerneInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionGay–Berne interaction.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
eps (
float) – Potential well depth.sig (
float) – Interaction range.cut (
float) – Cutoff distance of the interaction.k1 (
floatorstr) – Molecular elongation.k2 (
float, optional) – Ratio of the potential well depths for the side-by-side and end-to-end configurations.mu (
float, optional) – Adjustable exponent.nu (
float, optional) – Adjustable exponent.
- class espressomd.interactions.GenericLennardJonesInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionGeneralized Lennard-Jones potential.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
epsilon (
float) – Magnitude of the interaction.sigma (
float) – Interaction length scale.cutoff (
float) – Cutoff distance of the interaction.shift (
floatorstr{‘auto’}) – Constant shift of the potential. If'auto', a default value is computed from the other parameters. The LJ potential will be shifted by \(\epsilon\cdot\text{shift}\).offset (
float) – Offset distance of the interaction.e1 (
int) – Exponent of the repulsion term.e2 (
int) – Exponent of the attraction term.b1 (
float) – Prefactor of the repulsion term.b2 (
float) – Prefactor of the attraction term.delta (
float, optional) –LJGEN_SOFTCOREparameter delta. Allows control over how smoothly the potential drops to zero as lambda approaches zero.lam (
float, optional) –LJGEN_SOFTCOREparameter lambda. Tune the strength of the interaction.
- class espressomd.interactions.HarmonicBond(**kwargs)[source]¶
Bases:
BondedInteractionHarmonic bond.
- Parameters:
k (
float) – Magnitude of the bond interaction.r_0 (
float) – Equilibrium bond length.r_cut (
float, optional) – Maximum distance beyond which the bond is considered broken.
- class espressomd.interactions.HatInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionHat interaction.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
F_max (
float) – Magnitude of the interaction.cutoff (
float) – Cutoff distance of the interaction.
- class espressomd.interactions.HertzianInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionHertzian interaction.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
eps (
float) – Magnitude of the interaction.sig (
float) – Interaction length scale.
- class espressomd.interactions.IBM_Tribend(**kwargs)[source]¶
Bases:
BondedInteractionIBM Tribend bond.
See Figure C.2 in [Krüger, 2012].
- Parameters:
ind1, ind2, ind3, ind4 (
int) – First, second, third and fourth bonding partner. Used for initializing reference statekb (
float) – Bending modulusrefShape (
str, optional, {‘Flat’, ‘Initial’}) – Reference shape, default is'Flat'
- class espressomd.interactions.IBM_Triel(**kwargs)[source]¶
Bases:
BondedInteractionIBM Triel bond.
See Figure C.1 in [Krüger, 2012].
- Parameters:
ind1, ind2, ind3 (
int) – First, second and third bonding partner. Used for initializing reference statek1 (
float) – Shear elasticity for Skalak and Neo-Hookeank2 (
float, optional) – Area resistance for SkalakmaxDist (
float) – Gives an error if an edge becomes longer than maxDistelasticLaw (
str, {‘NeoHookean’, ‘Skalak’}) – Type of elastic bond
- class espressomd.interactions.IBM_VolCons(**kwargs)[source]¶
Bases:
BondedInteractionIBM volume conservation bond.
See Figure C.3 in [Krüger, 2012].
- Parameters:
softID (
int) – Used to identify the object to which this bond belongs. Each object (cell) needs its own ID. For performance reasons, it is best to start fromsoftID=0and increment by 1 for each subsequent bond.kappaV (
float) – Modulus for volume force
- class espressomd.interactions.LennardJonesCos2Interaction(**kwargs)[source]¶
Bases:
NonBondedInteractionSecond variant of the Lennard-Jones cosine interaction.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
epsilon (
float) – Magnitude of the interaction.sigma (
float) – Interaction length scale.offset (
float, optional) – Offset distance of the interaction.width (
float) – Width of interaction.
- property cutoff¶
- class espressomd.interactions.LennardJonesCosInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionLennard-Jones cosine interaction.
- set_params()¶
Set or update parameters for the interaction. Parameters marked as required become optional once the interaction has been activated for the first time; subsequent calls to this method update the existing values.
- Parameters:
epsilon (
float) – Magnitude of the interaction.sigma (
float) – Interaction length scale.cutoff (
float) – Cutoff distance of the interaction.offset (
float, optional) – Offset distance of the interaction.
- class espressomd.interactions.LennardJonesInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionStandard 6-12 Lennard-Jones potential.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
epsilon (
float) – Magnitude of the interaction.sigma (
float) – Interaction length scale.cutoff (
float) – Cutoff distance of the interaction.shift (
floatorstr{‘auto’}) – Constant shift of the potential. If'auto', a default value is computed fromsigmaandcutoff. The LJ potential will be shifted by \(4\epsilon\cdot\text{shift}\).offset (
float, optional) – Offset distance of the interaction.min (
float, optional) – Restricts the interaction to a minimal distance.
- class espressomd.interactions.MorseInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionMorse interaction.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
eps (
float) – The magnitude of the interaction.alpha (
float) – Stiffness of the Morse interaction.rmin (
float) – Distance of potential minimumcutoff (
float, optional) – Cutoff distance of the interaction.
- class espressomd.interactions.NonBondedInteraction(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperRepresents an instance of a non-bonded interaction, such as Lennard-Jones.
- deactivate()¶
Reset parameters for the interaction.
- class espressomd.interactions.NonBondedInteractionHandle(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperProvides access to all non-bonded interactions between two particle types.
- class espressomd.interactions.NonBondedInteractions(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperAccess to non-bonded interaction parameters via
[i,j], wherei, jare particle types. Returns aNonBondedInteractionHandleobject.- reset()¶
Reset all interaction parameters to their default values.
- class espressomd.interactions.OifGlobalForces(**kwargs)[source]¶
Bases:
BondedInteractionCharacterize the distribution of the force of the global mesh deformation onto individual vertices of the mesh.
Part of the Object-in-fluid method.
- Parameters:
A0_g (
float) – Relaxed area of the meshka_g (
float) – Area coefficientV0 (
float) – Relaxed volume of the meshkv (
float) – Volume coefficient
- class espressomd.interactions.OifLocalForces(**kwargs)[source]¶
Bases:
BondedInteractionCharacterize the deformation of two triangles sharing an edge.
Part of the Object-in-fluid method.
- Parameters:
r0 (
float) – Equilibrium bond length of triangle edgesks (
float) – Non-linear stretching coefficient of triangle edgeskslin (
float) – Linear stretching coefficient of triangle edgesphi0 (
float) – Equilibrium angle between the two triangleskb (
float) – Bending coefficient for the angle between the two trianglesA01 (
float) – Equilibrium surface of the first triangleA02 (
float) – Equilibrium surface of the second trianglekal (
float) – Stretching coefficient of a triangle surfacekvisc (
float) – Viscous coefficient of the triangle vertices
- class espressomd.interactions.QuarticBond(**kwargs)[source]¶
Bases:
BondedInteractionQuartic bond.
- Parameters:
k0 (
float) – Magnitude of the square term.k1 (
float) – Magnitude of the fourth order term.r (
float) – Equilibrium bond length.r_cut (
float) – Maximum interaction length.
- class espressomd.interactions.RigidBond(**kwargs)[source]¶
Bases:
BondedInteractionRigid bond.
- Parameters:
r (
float) – Bond length.ptol (
float, optional) – Tolerance for positional deviations.vtop (
float, optional) – Tolerance for velocity deviations.
- class espressomd.interactions.SmoothStepInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionSmooth step interaction.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
d (
float) – Short range repulsion parameter.n (
int, optional) – Exponent of short range repulsion.eps (
float) – Magnitude of the second (soft) repulsion.k0 (
float, optional) – Exponential factor in second (soft) repulsion.sig (
float, optional) – Length scale of second (soft) repulsion.cutoff (
float) – Cutoff distance of the interaction.
- class espressomd.interactions.SoftSphereInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionSoft sphere interaction.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
a (
float) – Magnitude of the interaction.n (
float) – Exponent of the power law.cutoff (
float) – Cutoff distance of the interaction.offset (
float, optional) – Offset distance of the interaction.
- class espressomd.interactions.TabulatedAngle(*args, **kwargs)[source]¶
Bases:
BondedInteractionTabulated bond angle.
- Parameters:
energy (array_like of
float) – The energy table for the range \(0-\pi\).force (array_like of
float) – The force table for the range \(0-\pi\).
- pi = 3.14159265358979¶
- class espressomd.interactions.TabulatedDihedral(*args, **kwargs)[source]¶
Bases:
BondedInteractionTabulated bond dihedral.
- Parameters:
energy (array_like of
float) – The energy table for the range \(0-2\pi\).force (array_like of
float) – The force table for the range \(0-2\pi\).
- pi = 3.14159265358979¶
- class espressomd.interactions.TabulatedDistance(**kwargs)[source]¶
Bases:
BondedInteractionTabulated bond length.
- Parameters:
min (
float) – The minimal interaction distance.max (
float) – The maximal interaction distance.energy (array_like of
float) – The energy table.force (array_like of
float) – The force table.
- class espressomd.interactions.TabulatedNonBonded(**kwargs)[source]¶
Bases:
NonBondedInteractionTabulated interaction.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
min (
float,) – The minimal interaction distance.max (
float,) – The maximal interaction distance.energy (array_like of
float) – The energy table.force (array_like of
float) – The force table.
- property cutoff¶
- class espressomd.interactions.ThermalizedBond(**kwargs)[source]¶
Bases:
BondedInteractionThermalized bond.
- Parameters:
temp_com (
float) – Temperature of the Langevin thermostat for the center of mass of the particle pair.gamma_com (
float) – Friction coefficient of the Langevin thermostat for the center of mass of the particle pair.temp_distance (
float) – Temperature of the Langevin thermostat for the distance vector of the particle pair.gamma_distance (
float) – Friction coefficient of the Langevin thermostat for the distance vector of the particle pair.r_cut (
float, optional) – Maximum distance beyond which the bond is considered broken.
- class espressomd.interactions.TholeInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionThole interaction.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
scaling_coeff (
float) – The factor used in the Thole damping function between polarizable particles i and j. Usually calculated by the polarizabilities \(\alpha_i\), \(\alpha_j\) and damping parameters \(a_i\), \(a_j\) via \(s_{ij} = \frac{(a_i+a_j)/2}{((\alpha_i\cdot\alpha_j)^{1/2})^{1/3}}\)q1q2 (
float) – Charge factor of the involved charges. Has to be set because it acts only on the portion of the Drude core charge that is associated to the dipole of the atom. For charged, polarizable atoms that charge is not equal to the particle charge property.
- class espressomd.interactions.Virtual(**kwargs)[source]¶
Bases:
BondedInteractionVirtual bond.
- class espressomd.interactions.WCAInteraction(**kwargs)[source]¶
Bases:
NonBondedInteractionStandard 6-12 Weeks-Chandler-Andersen potential.
- set_params()¶
Set new parameters for the interaction.
- Parameters:
epsilon (
float) – Magnitude of the interaction.sigma (
float) – Interaction length scale.
- property cutoff¶
espressomd.lb module¶
- class espressomd.lb.HydrodynamicInteraction(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperBase class for LB implementations.
- mach_limit()[source]¶
The fluid velocity is limited to \(v_{\mathrm{max}} = 0.20\) (see quasi-incompressible limit in [Krüger et al., 2017], chapter 7, page 272), which corresponds to Mach 0.35.
The relative error in the fluid density between a compressible fluid and an incompressible fluid at Mach 0.30 is less than 5% (see constant density assumption in [Kundu et al., 2001] chapter 16, page 663). Since the speed of sound is \(c_s = 1 / \sqrt{3}\) in LB velocity units in a D3Q19 lattice, the velocity limit at Mach 0.30 is \(v_{\mathrm{max}} = 0.30 / \sqrt{3} \approx 0.17\). At Mach 0.35 the relative error is around 6% and \(v_{\mathrm{max}} = 0.35 / \sqrt{3} \approx 0.20\).
- Returns:
v_max – The Mach limit expressed in LB velocity units.
- Return type:
float
- property pressure_tensor¶
- class espressomd.lb.LBFluidNodeWalberla(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper- property boundary¶
returns: *
VelocityBounceBack– If the node is a boundary node * None – If the node is not a boundary node
- property boundary_force¶
- property density¶
- property index¶
- property is_boundary¶
- property last_applied_force¶
- property population¶
- property pressure_tensor¶
- property pressure_tensor_neq¶
- property velocity¶
- class espressomd.lb.LBFluidSliceWalberla(*args, **kwargs)[source]¶
Bases:
ScriptInterfaceHelper- property boundary¶
returns: * (N, M, L) array_like of
VelocityBounceBack– If the nodes are boundary nodes * (N, M, L) array_like ofNone– If the nodes are not boundary nodes
- property boundary_force¶
- property density¶
- property is_boundary¶
- property last_applied_force¶
- property population¶
- property pressure_tensor¶
- property pressure_tensor_neq¶
- property velocity¶
- class espressomd.lb.LBFluidWalberla(*args, **kwargs)[source]¶
Bases:
HydrodynamicInteraction,LatticeModelThe lattice-Boltzmann method for hydrodynamics using waLBerla. If argument
latticeis not provided, one will be default constructed if an argumentagridis provided.- Parameters:
lattice (
espressomd.lb.LatticeWalberla) – Lattice object. If not provided, a default one will be constructed using theagridparameter.agrid (
float) – Lattice constant. The box size in every direction must be an integer multiple ofagrid. Cannot be provided together withlattice.tau (
float) – LB time step, must be an integer multiple of the MD time step.density (
float) – Fluid density.kinematic_viscosity (
float) – Fluid kinematic viscosity.ext_force_density ((3,) array_like of
float, optional) – Force density applied on the fluid.kT (
float, optional) – Thermal energy of the simulated heat bath (for thermalized fluids). Set it to 0 for an unthermalized fluid.seed (
int, optional) – Initial counter value (or seed) of the philox RNG. Required for a thermalized fluid. Must be positive.single_precision (
bool, optional) – Use single-precision floating-point arithmetic.
- get_interpolated_velocity()¶
Get LB fluid velocity at specified position.
- Parameters:
pos ((3,) array_like of
float) – The position at which velocity is requested.- Returns:
v – The LB fluid velocity at
pos.- Return type:
(3,) array_like
float
- add_force_at_pos():
Adds a force to the fluid at given position.
- Parameters:
pos ((3,) array_like of
float) – The position at which the force will be added.force ((3,) array_like of
float) – The force vector which will be distributed at the position.
- clear_boundaries()¶
Remove velocity bounce-back boundary conditions.
- save_checkpoint()¶
Write LB node populations and boundary conditions to a file.
- Parameters:
path (
str) – Destination file path.binary (
bool) – Whether to write in binary or ASCII mode.
- load_checkpoint()¶
Load LB node populations and boundary conditions from a file.
- Parameters:
path (
str) – File path to read from.binary (
bool) – Whether to read in binary or ASCII mode.
- add_vtk_writer()¶
Attach a VTK writer.
- Parameters:
vtk (
espressomd.lb.VTKOutput) – VTK writer.
- remove_vtk_writer()¶
Detach a VTK writer.
- Parameters:
vtk (
espressomd.lb.VTKOutput) – VTK writer.
- clear_vtk_writers()¶
Detach all VTK writers.
- add_boundary_from_shape(shape, velocity=array([0., 0., 0.]), boundary_type=<class 'espressomd.lb.VelocityBounceBack'>)[source]¶
Set velocity bounce-back boundary conditions from a shape.
- Parameters:
shape (
espressomd.shapes.Shape) – Shape to rasterize.velocity ((3,) or (L, M, N, 3) array_like of
float, optional) – Slip velocity. By default no-slip boundary conditions are used. If a vector of 3 values, a uniform slip velocity is used, otherwiseL, M, Nmust be equal to the LB grid dimensions.boundary_type (Union[
VelocityBounceBack] (optional)) – Type of the boundary condition.
- class espressomd.lb.LBFluidWalberlaGPU(*args, **kwargs)[source]¶
Bases:
HydrodynamicInteractionInitialize the lattice-Boltzmann method for hydrodynamic flow using waLBerla for the GPU. See
HydrodynamicInteractionfor the list of parameters.
- class espressomd.lb.VTKOutput(*args, **kwargs)[source]¶
Bases:
VTKOutputBaseCreate a VTK writer.
Files are written to
<base_folder>/<identifier>/<prefix>_*.vtu. Summary is written to<base_folder>/<identifier>.pvd.Manual VTK callbacks can be called at any time to take a snapshot of the current state of the LB fluid.
Automatic VTK callbacks can be disabled at any time and re-enabled later. Please note that the internal VTK counter is no longer incremented when an automatic callback is disabled, which means the number of LB steps between two frames will not always be an integer multiple of
delta_N.- Parameters:
identifier (
str) – Name of the VTK writer.observables (
list, {‘density’, ‘velocity_vector’, ‘pressure_tensor’}) – List of observables to write to the VTK files.delta_N (
int) – Write frequency. If this value is 0 (default), the object is a manual VTK callback that must be triggered manually. Otherwise, it is an automatic callback that is added to the time loop and writes everydelta_NLB steps.base_folder (
str(optional), default is ‘vtk_out’) – Path to the output VTK folder.prefix (
str(optional), default is ‘simulation_step’) – Prefix for VTK files.
- class espressomd.lb.VelocityBounceBack(velocity)[source]¶
Bases:
objectHold velocity information for the velocity bounce back boundary condition at a single node.
- espressomd.lb.calc_cylinder_tangential_vectors(center, agrid, offset, node_indices)[source]¶
Utility function to calculate a constant slip velocity tangential to the surface of a cylinder.
- Parameters:
center ((3,) array_like of
float) – Center of the cylinder.agrid (
float) – LB agrid.offset (
float) – LB offset.node_indices ((N, 3) array_like of
int) – Indices of the boundary surface nodes.
- Returns:
The unit vectors tangential to the surface of a cylinder.
- Return type:
(N, 3) array_like of
float
- espressomd.lb.edge_detection(boundary_mask, periodicity)[source]¶
Find boundary nodes in contact with the fluid. Relies on a convolution kernel constructed from the D3Q19 stencil.
- Parameters:
boundary_mask ((N, M, L) array_like of
bool) – Bitmask for the rasterized boundary geometry.periodicity ((3,) array_like of
bool) – Bitmask for the box periodicity.
- Returns:
The indices of the boundary nodes at the interface with the fluid.
- Return type:
(N, 3) array_like of
int
espressomd.lees_edwards module¶
- class espressomd.lees_edwards.LeesEdwards(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperInterface to the Lees–Edwards boundary conditions. When writing H5MD files, the shear direction and shear plane normals are written as integers instead of characters, with 0 = x-axis, 1 = y-axis, 2 = z-axis.
- protocol¶
Lees–Edwards protocol.
- Type:
object
- shear_velocity¶
Current shear velocity.
- Type:
float
- pos_offset¶
Current position offset
- Type:
float
- shear_direction¶
Shear direction.
- Type:
str, {‘x’, ‘y’, ‘z’}
- shear_plane_normal¶
Shear plane normal.
- Type:
str, {‘x’, ‘y’, ‘z’}
- set_boundary_conditions()¶
Set a protocol, the shear direction and shear normal.
- Parameters:
protocol (
object)shear_direction (
str, {‘x’, ‘y’, ‘z’})shear_plane_normal (
str, {‘x’, ‘y’, ‘z’})
- class espressomd.lees_edwards.LinearShear(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperLees–Edwards protocol for linear shear.
- Parameters:
initial_pos_offset (
float) – Positional offset at the Lees–Edwards boundary at t=0.shear_velocity (
float) – Shear velocity (velocity jump) across the Lees–Edwards boundary.
- class espressomd.lees_edwards.Off(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperLees–Edwards protocol resulting in un-shifted boundaries.
- class espressomd.lees_edwards.OscillatoryShear(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperLees–Edwards protocol for oscillatory shear.
- Parameters:
initial_pos_offset (
float) – Positional offset at the Lees–Edwards boundary at t=0.amplitude (
float) – Maximum amplitude of the positional offset at the Lees–Edwards boundary.omega (
float) – Radian frequency of the oscillation.time_0 (
float) – Time offset of the oscillation.
espressomd.magnetostatics module¶
- class espressomd.magnetostatics.Container(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper
- class espressomd.magnetostatics.DLC(**kwargs)[source]¶
Bases:
MagnetostaticInteractionElectrostatics solver for systems with two periodic dimensions. See Dipolar Layer Correction (DLC) for more details.
Notes
At present, the empty gap (volume without any particles), is assumed to be along the z-axis. As a reference for the DLC method, see [Bródka, 2004].
- Parameters:
actor (object derived of
MagnetostaticInteraction, required) – Base solver.gap_size (
float) – The gap size gives the height \(h\) of the empty region between the system box and the neighboring artificial images. ESPResSo checks that the gap is empty and will throw an error if it isn’t. Therefore you should really make sure that the gap region is empty (e.g. with wall constraints).maxPWerror (
float) – Maximal pairwise error of the potential and force.far_cut (
float, optional) – Cutoff of the exponential sum.
- class espressomd.magnetostatics.DipolarBarnesHutGpu(**kwargs)[source]¶
Bases:
MagnetostaticInteractionCalculates magnetostatic interactions by direct summation over all pairs. See Barnes-Hut octree sum on GPU for more details.
TODO: If the system has periodic boundaries, the minimum image convention is applied.
Requires feature
DIPOLAR_BARNES_HUT, which depends onDIPOLESandCUDA.- Parameters:
epssq (
float, optional) – Squared skin of the octant cells.itolsq (
float, optional) – Squared inverse fraction of the octant cells.
- class espressomd.magnetostatics.DipolarDirectSumCpu(**kwargs)[source]¶
Bases:
MagnetostaticInteractionCalculate magnetostatic interactions by direct summation over all pairs. See Dipolar direct sum for more details.
If the system has periodic boundaries, the minimum image convention is applied in the respective directions when no replicas are used. When replicas are used,
n_replicascopies of the system are taken into account in the respective directions, and a spherical cutoff is applied.- Parameters:
prefactor (
float) – Magnetostatics prefactor (\(\mu_0/(4\pi)\))n_replicas (
int, optional) – Number of replicas to be taken into account at periodic boundaries.
- class espressomd.magnetostatics.DipolarDirectSumGpu(**kwargs)[source]¶
Bases:
MagnetostaticInteractionCalculate magnetostatic interactions by direct summation over all pairs. See Dipolar direct sum for more details.
If the system has periodic boundaries, the minimum image convention is applied in the respective directions.
This is the GPU version of
espressomd.magnetostatics.DipolarDirectSumCpubut uses floating point precision.Requires feature
DIPOLAR_DIRECT_SUM, which depends onDIPOLESandCUDA.- Parameters:
prefactor (
float) – Magnetostatics prefactor (\(\mu_0/(4\pi)\))
- class espressomd.magnetostatics.DipolarP3M(**kwargs)[source]¶
Bases:
MagnetostaticInteractionCalculate magnetostatic interactions using the dipolar P3M method. See Dipolar P3M for more details.
- Parameters:
prefactor (
float) – Magnetostatics prefactor (\(\mu_0/(4\pi)\))accuracy (
float) – P3M tunes its parameters to provide this target accuracy.alpha (
float) – Ewald parameter.cao (
int) – Charge-assignment order, an integer between 1 and 7.mesh (
intor (3,) array_like ofint) – The number of mesh points in x, y and z direction. Use a single value for cubic boxes.mesh_off ((3,) array_like of
float, optional) – Mesh offset.r_cut (
float) – Real space cutoff.tune (
bool, optional) – Activate/deactivate the tuning method on activation (default isTrue, i.e., activated).timings (
int) – Number of force calculations during tuning.
- class espressomd.magnetostatics.MagnetostaticInteraction(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperCommon interface for magnetostatics solvers.
- Parameters:
prefactor (
float) – Magnetostatics prefactor \(\frac{\mu_0\mu}{4\pi}\)
- class espressomd.magnetostatics.Scafacos(**kwargs)[source]¶
Bases:
MagnetostaticInteractionCalculate the dipolar interaction using dipoles-capable methods from the ScaFaCoS library. See ScaFaCoS magnetostatics for more details.
- Parameters:
prefactor (
float) – Magnetostatics prefactor (\(\mu_0/(4\pi)\)).method_name (
str) – Name of the ScaFaCoS method to use.method_params (
dict) – Dictionary with the key-value pairs of the method parameters as defined in ScaFaCoS. Note that the values are cast to strings to match ScaFaCoS’ interface.
- get_available_methods()¶
List long-range methods available in the ScaFaCoS library.
espressomd.math module¶
- class espressomd.math.CylindricalTransformationParameters(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperClass to hold and validate the parameters needed for a cylindrical transformation. The three parameters are available as attributes but are read-only.
- Parameters:
center ((3,) array_like of
float, default = [0, 0, 0]) – Position of the origin of the cylindrical coordinate system.axis ((3,) array_like of
float, default = [0, 0, 1]) – Orientation vector of thez-axis of the cylindrical coordinate system.orientation ((3,) array_like of
float, default = [1, 0, 0]) – The axis on whichphi = 0.
Notes
If you provide no arguments, the defaults above are set. If you provide only a
centerand anaxis, anorientationwill be automatically generated that is orthogonal toaxis.
- espressomd.math.calc_quaternions_from_angles(polar_angle, azimuthal_angle)[source]¶
Convert the spherical coordinate representation of a vector to a quaternion that corresponds to that direction. Note that this quaternion is not unique, as the third degree of freedom (rotation around the given vector) is not specified.
- Parameters:
polar_angle (
float) – The polar angle (angle with the z-axis) in the range [0, pi].azimuthal_angle (
float) – The azimuthal angle (angle with the x-axis) in the range [0, 2 pi).
- Returns:
quaternion
- Return type:
(4,) list of
float
espressomd.observables module¶
- class espressomd.observables.BondAngles(**kwargs)[source]¶
Bases:
ObservableCalculates the angles between bonds of particles with given ids along a polymer chain.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(N - 2,)
ndarrayoffloat
- class espressomd.observables.BondDihedrals(**kwargs)[source]¶
Bases:
ObservableCalculates the dihedrals between particles with given ids along a polymer chain.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(N - 3,)
ndarrayoffloat
- class espressomd.observables.ComPosition(**kwargs)[source]¶
Bases:
ObservableCalculates the center of mass for particles with given ids.
Note that virtual sites are not included since they do not have a meaningful mass.
Output format: \(\frac{1}{\sum_i m_i} \left( \sum_i m_i r^x_i, \sum_i m_i r^y_i, \sum_i m_i r^z_i\right)\)
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(3,)
ndarrayoffloat
- class espressomd.observables.ComVelocity(**kwargs)[source]¶
Bases:
ObservableCalculates the center of mass velocity for particles with given ids.
Note that virtual sites are not included since they do not have a meaningful mass.
Output format: \(\frac{1}{\sum_i m_i} \left( \sum_i m_i v^x_i, \sum_i m_i v^y_i, \sum_i m_i v^z_i\right)\)
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(3,)
ndarrayoffloat
- class espressomd.observables.CosPersistenceAngles(**kwargs)[source]¶
Bases:
ObservableCalculates the cosine of mutual bond angles for chained particles with given ids.
The i-th value of the result contains the cosine of the angle between bonds that are separated by i bonds. The values are averaged over the chain.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(N - 2,)
ndarrayoffloat
- class espressomd.observables.CylindricalDensityProfile(transform_params=<espressomd.math.CylindricalTransformationParameters object>, **kwargs)[source]¶
Bases:
CylindricalProfileObservableCalculates the particle density in cylindrical coordinates.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.transform_params (
espressomd.math.CylindricalTransformationParameters, optional) – Parameters of the cylinder transformation. Defaults to the default ofespressomd.math.CylindricalTransformationParametersn_r_bins (
int, default = 1) – Number of bins in radial direction.n_phi_bins (
int, default = 1) – Number of bins for the azimuthal direction.n_z_bins (
int, default = 1) – Number of bins inzdirection.min_r (
float, default = 0) – Minimumrto consider.min_phi (
float, default = \(-\pi\)) – Minimumphito consider. Must be in \([-\pi,\pi)\).min_z (
float) – Minimumzto consider.max_r (
float) – Maximumrto consider.max_phi (
float, default = \(\pi\)) – Maximumphito consider. Must be in \((-\pi,\pi]\).max_z (
float) – Maximumzto consider.
- calculate()¶
Run the observable.
- Return type:
(
n_r_bins,n_phi_bins,n_z_bins)ndarrayoffloat
- class espressomd.observables.CylindricalFluxDensityProfile(transform_params=<espressomd.math.CylindricalTransformationParameters object>, **kwargs)[source]¶
Bases:
CylindricalProfileObservableCalculates the particle flux density in cylindrical coordinates.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.transform_params (
espressomd.math.CylindricalTransformationParameters, optional) – Parameters of the cylinder transformation. Defaults to the default ofespressomd.math.CylindricalTransformationParametersn_r_bins (
int, default = 1) – Number of bins in radial direction.n_phi_bins (
int, default = 1) – Number of bins for the azimuthal direction.n_z_bins (
int, default = 1) – Number of bins inzdirection.min_r (
float, default = 0) – Minimumrto consider.min_phi (
float, default = \(-\pi\)) – Minimumphito consider. Must be in \([-\pi,\pi)\).min_z (
float) – Minimumzto consider.max_r (
float) – Maximumrto consider.max_phi (
float, default = \(\pi\)) – Maximumphito consider. Must be in \((-\pi,\pi]\).max_z (
float) – Maximumzto consider.
- calculate()¶
Run the observable.
- Returns:
The fourth dimension of the array stores the histogram for the radial distance, azimuth and axial coordinate of the particle flux density field, respectively.
- Return type:
(
n_r_bins,n_phi_bins,n_z_bins, 3)ndarrayoffloat
- class espressomd.observables.CylindricalLBFluxDensityProfileAtParticlePositions(transform_params=<espressomd.math.CylindricalTransformationParameters object>, **kwargs)[source]¶
Bases:
CylindricalProfileObservableCalculates the LB fluid flux density at the particle positions in cylindrical coordinates.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.transform_params (
espressomd.math.CylindricalTransformationParameters, optional) – Parameters of the cylinder transformation. Defaults to the default ofespressomd.math.CylindricalTransformationParametersn_r_bins (
int, default = 1) – Number of bins in radial direction.n_phi_bins (
int, default = 1) – Number of bins for the azimuthal direction.n_z_bins (
int, default = 1) – Number of bins inzdirection.min_r (
float, default = 0) – Minimumrto consider.min_phi (
float, default = \(-\pi\)) – Minimumphito consider. Must be in \([-\pi,\pi)\).min_z (
float) – Minimumzto consider.max_r (
float) – Maximumrto consider.max_phi (
float, default = \(\pi\)) – Maximumphito consider. Must be in \((-\pi,\pi]\).max_z (
float) – Maximumzto consider.
- calculate()¶
Run the observable.
- Returns:
The fourth dimension of the array stores the histogram for the radial distance, azimuth and axial coordinate of the LB flux density field, respectively.
- Return type:
(
n_r_bins,n_phi_bins,n_z_bins, 3)ndarrayoffloat
- class espressomd.observables.CylindricalLBVelocityProfile(transform_params=<espressomd.math.CylindricalTransformationParameters object>, **kwargs)[source]¶
Bases:
CylindricalProfileObservableCalculates the LB fluid velocity profile in cylindrical coordinates.
This observable samples the fluid in on a regular grid defined by variable
sampling_density. Note that a small delta leads to a large number of sample points and carries a performance cost.- Parameters:
transform_params (
espressomd.math.CylindricalTransformationParameters, optional) – Parameters of the cylinder transformation. Defaults to the default ofespressomd.math.CylindricalTransformationParametersn_r_bins (
int, default = 1) – Number of bins in radial direction.n_phi_bins (
int, default = 1) – Number of bins for the azimuthal direction.n_z_bins (
int, default = 1) – Number of bins inzdirection.min_r (
float, default = 0) – Minimumrto consider.min_phi (
float, default = \(-\pi\)) – Minimumphito consider. Must be in \([-\pi,\pi)\).min_z (
float) – Minimumzto consider.max_r (
float) – Maximumrto consider.max_phi (
float, default = \(\pi\)) – Maximumphito consider. Must be in \((-\pi,\pi]\).max_z (
float) – Maximumzto consider.sampling_density (
float) – Samples per unit volume for the LB velocity interpolation.
- calculate()¶
Run the observable.
- Returns:
The fourth dimension of the array stores the histogram for the radial distance, azimuth and axial coordinate of the LB velocity field, respectively.
- Return type:
(
n_r_bins,n_phi_bins,n_z_bins, 3)ndarrayoffloat
- class espressomd.observables.CylindricalLBVelocityProfileAtParticlePositions(transform_params=<espressomd.math.CylindricalTransformationParameters object>, **kwargs)[source]¶
Bases:
CylindricalProfileObservableCalculates the LB fluid velocity at the particle positions in cylindrical coordinates.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.transform_params (
espressomd.math.CylindricalTransformationParameters, optional) – Parameters of the cylinder transformation. Defaults to the default ofespressomd.math.CylindricalTransformationParametersn_r_bins (
int, default = 1) – Number of bins in radial direction.n_phi_bins (
int, default = 1) – Number of bins for the azimuthal direction.n_z_bins (
int, default = 1) – Number of bins inzdirection.min_r (
float, default = 0) – Minimumrto consider.min_phi (
float, default = \(-\pi\)) – Minimumphito consider. Must be in \([-\pi,\pi)\).min_z (
float) – Minimumzto consider.max_r (
float) – Maximumrto consider.max_phi (
float, default = \(\pi\)) – Maximumphito consider. Must be in \((-\pi,\pi]\).max_z (
float) – Maximumzto consider.
- calculate()¶
Run the observable.
- Returns:
The fourth dimension of the array stores the histogram for the radial distance, azimuth and axial coordinate of the LB velocity field, respectively.
- Return type:
(
n_r_bins,n_phi_bins,n_z_bins, 3)ndarrayoffloat
- class espressomd.observables.CylindricalProfileObservable(transform_params=<espressomd.math.CylindricalTransformationParameters object>, **kwargs)[source]¶
Bases:
ProfileObservableBase class for observables that work with cylinder coordinates
- class espressomd.observables.CylindricalVelocityProfile(transform_params=<espressomd.math.CylindricalTransformationParameters object>, **kwargs)[source]¶
Bases:
CylindricalProfileObservableCalculates the particle velocity profile in cylindrical coordinates.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.transform_params (
espressomd.math.CylindricalTransformationParameters, optional) – Parameters of the cylinder transformation. Defaults to the default ofespressomd.math.CylindricalTransformationParametersn_r_bins (
int, default = 1) – Number of bins in radial direction.n_phi_bins (
int, default = 1) – Number of bins for the azimuthal direction.n_z_bins (
int, default = 1) – Number of bins inzdirection.min_r (
float, default = 0) – Minimumrto consider.min_phi (
float, default = \(-\pi\)) – Minimumphito consider. Must be in \([-\pi,\pi)\).min_z (
float) – Minimumzto consider.max_r (
float) – Maximumrto consider.max_phi (
float, default = \(\pi\)) – Maximumphito consider. Must be in \((-\pi,\pi]\).max_z (
float) – Maximumzto consider.
- calculate()¶
Run the observable.
- Returns:
The fourth dimension of the array stores the histogram for the radial distance, azimuth and axial coordinate of the particle velocity field, respectively.
- Return type:
(
n_r_bins,n_phi_bins,n_z_bins, 3)ndarrayoffloat
- class espressomd.observables.DPDStress(**kwargs)[source]¶
Bases:
ObservableCalculates the non-equilibrium contribution of the DPD interaction to the stress tensor.
- Parameters:
None
- calculate()¶
Run the observable.
- Return type:
(3, 3)
ndarrayoffloat
- class espressomd.observables.DensityProfile(**kwargs)[source]¶
Bases:
ProfileObservableCalculates the particle density profile for particles with given ids.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.n_x_bins (
int) – Number of bins inxdirection.n_y_bins (
int) – Number of bins inydirection.n_z_bins (
int) – Number of bins inzdirection.min_x (
float) – Minimumxto consider.min_y (
float) – Minimumyto consider.min_z (
float) – Minimumzto consider.max_x (
float) – Maximumxto consider.max_y (
float) – Maximumyto consider.max_z (
float) – Maximumzto consider.
- calculate()¶
Run the observable.
- Return type:
(
n_x_bins,n_y_bins,n_z_bins)ndarrayoffloat
- class espressomd.observables.DipoleMoment(**kwargs)[source]¶
Bases:
ObservableCalculates the electric dipole moment for particles with given ids.
Output format: \(\left(\sum_i q_i r^x_i, \sum_i q_i r^y_i, \sum_i q_i r^z_i\right)\)
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(3,)
ndarrayoffloat
- class espressomd.observables.Energy(**kwargs)[source]¶
Bases:
ObservableCalculates the total energy.
- calculate()¶
Run the observable.
- Return type:
float
- class espressomd.observables.FluxDensityProfile(**kwargs)[source]¶
Bases:
ProfileObservableCalculates the particle flux density for particles with given ids.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.n_x_bins (
int) – Number of bins inxdirection.n_y_bins (
int) – Number of bins inydirection.n_z_bins (
int) – Number of bins inzdirection.min_x (
float) – Minimumxto consider.min_y (
float) – Minimumyto consider.min_z (
float) – Minimumzto consider.max_x (
float) – Maximumxto consider.max_y (
float) – Maximumyto consider.max_z (
float) – Maximumzto consider.
- calculate()¶
Run the observable.
- Returns:
The fourth dimension of the array stores the histogram for the x, y and z components of the flux density, respectively.
- Return type:
(
n_x_bins,n_y_bins,n_z_bins, 3)ndarrayoffloat
- class espressomd.observables.ForceDensityProfile(**kwargs)[source]¶
Bases:
ProfileObservableCalculates the force density profile for particles with given ids.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.n_x_bins (
int) – Number of bins inxdirection.n_y_bins (
int) – Number of bins inydirection.n_z_bins (
int) – Number of bins inzdirection.min_x (
float) – Minimumxto consider.min_y (
float) – Minimumyto consider.min_z (
float) – Minimumzto consider.max_x (
float) – Maximumxto consider.max_y (
float) – Maximumyto consider.max_z (
float) – Maximumzto consider.
- calculate()¶
Run the observable.
- Returns:
The fourth dimension of the array stores the histogram for the x, y and z components of the force, respectively.
- Return type:
(
n_x_bins,n_y_bins,n_z_bins, 3)ndarrayoffloat
- class espressomd.observables.LBFluidPressureTensor(**kwargs)[source]¶
Bases:
ObservableCalculates the average pressure tensor of the LB fluid for all nodes.
- Parameters:
None
- calculate()¶
Run the observable.
- Return type:
(3, 3)
ndarrayoffloat
- class espressomd.observables.LBVelocityProfile(**kwargs)[source]¶
Bases:
ProfileObservableCalculates the LB fluid velocity profile.
This observable samples the fluid in on a regular grid defined by the variables
sampling_*. Note that a small delta leads to a large number of sample points and carries a performance cost.- Parameters:
n_x_bins (
int) – Number of bins inxdirection.n_y_bins (
int) – Number of bins inydirection.n_z_bins (
int) – Number of bins inzdirection.min_x (
float) – Minimumxto consider.min_y (
float) – Minimumyto consider.min_z (
float) – Minimumzto consider.max_x (
float) – Maximumxto consider.max_y (
float) – Maximumyto consider.max_z (
float) – Maximumzto consider.sampling_delta_x (
float, default=1.0) – Spacing for the sampling grid inx-direction.sampling_delta_y (
float, default=1.0) – Spacing for the sampling grid iny-direction.sampling_delta_z (
float, default=1.0) – Spacing for the sampling grid inz-direction.sampling_offset_x (
float, default=0.0) – Offset for the sampling grid inx-direction.sampling_offset_y (
float, default=0.0) – Offset for the sampling grid iny-direction.sampling_offset_z (
float, default=0.0) – Offset for the sampling grid inz-direction.allow_empty_bins (
bool, default=False) – Whether or not to allow bins that will not be sampled at all.
- calculate()¶
Run the observable.
- Returns:
The fourth dimension of the array stores the histogram for the x, y and z components of the LB velocity, respectively.
- Return type:
(
n_x_bins,n_y_bins,n_z_bins, 3)ndarrayoffloat
- class espressomd.observables.MagneticDipoleMoment(**kwargs)[source]¶
Bases:
ObservableCalculates the magnetic dipole moment for particles with given ids.
Output format: \(\left(\sum_i \mu^x_i, \sum_i \mu^y_i, \sum_i \mu^z_i\right)\)
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(3,)
ndarrayoffloat
- class espressomd.observables.Observable(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperBase class for all observables.
- shape()¶
Get the shape of the numpy array returned by the observable.
- class espressomd.observables.ParticleAngularVelocities(**kwargs)[source]¶
Bases:
ObservableCalculates the angular velocity (omega) in the spaced-fixed frame of reference
Output format: \((\omega^x_1,\ \omega^y_1,\ \omega^z_1),\ (\omega^x_2,\ \omega^y_2,\ \omega^z_2), \dots,\ (\omega^x_n,\ \omega^y_n,\ \omega^z_n)\).
The particles are ordered according to the list of ids passed to the observable.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(N, 3)
ndarrayoffloat
- class espressomd.observables.ParticleBodyAngularVelocities(**kwargs)[source]¶
Bases:
ObservableCalculates the angular velocity (omega) in the particles’ body-fixed frame of reference.
For each particle, the body-fixed frame of reference is obtained from the particle’s orientation stored in the quaternions.
Output format: \((\omega^x_1,\ \omega^y_1,\ \omega^z_1),\ (\omega^x_2,\ \omega^y_2,\ \omega^z_2), \dots,\ (\omega^x_n,\ \omega^y_n,\ \omega^z_n)\).
The particles are ordered according to the list of ids passed to the observable.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(N, 3)
ndarrayoffloat
- class espressomd.observables.ParticleBodyVelocities(**kwargs)[source]¶
Bases:
ObservableCalculates the particle velocity in the particles’ body-fixed frame of reference.
For each particle, the body-fixed frame of reference is obtained from the particle’s orientation stored in the quaternions.
Output format: \((v^x_1,\ v^y_1,\ v^z_1),\ (v^x_2,\ v^y_2,\ v^z_2),\ \dots,\ (v^x_n,\ v^y_n,\ v^z_n)\).
The particles are ordered according to the list of ids passed to the observable.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(N, 3)
ndarrayoffloat
- class espressomd.observables.ParticleDipoleFields(**kwargs)[source]¶
Bases:
ObservableCalculates the particle dipole fields for particles with given ids.
Output format: \((h^x_1,\ h^y_1,\ h^z_1),\ (h^x_2,\ h^y_2,\ h^z_2),\ \dots,\ (h^x_n,\ h^y_n,\ h^z_n)\).
The particles are ordered according to the list of ids passed to the observable.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(N, 3)
ndarrayoffloat
- class espressomd.observables.ParticleDirectors(**kwargs)[source]¶
Bases:
ObservableCalculates the particle directors for particles with given ids.
Output format: \((d^x_1,\ d^y_1,\ d^z_1),\ (d^x_2,\ d^y_2,\ d^z_2),\ \dots,\ (d^x_n,\ d^y_n,\ d^z_n)\).
The particles are ordered according to the list of ids passed to the observable.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(N, 3)
ndarrayoffloat
- class espressomd.observables.ParticleDistances(**kwargs)[source]¶
Bases:
ObservableCalculates the distances between particles with given ids along a polymer chain.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(N - 1,)
ndarrayoffloat
- class espressomd.observables.ParticleForces(**kwargs)[source]¶
Bases:
ObservableCalculates the particle forces for particles with given ids.
Output format: \((f^x_1,\ f^y_1,\ f^z_1),\ (f^x_2,\ f^y_2,\ f^z_2),\ \dots,\ (f^x_n,\ f^y_n,\ f^z_n)\).
The particles are ordered according to the list of ids passed to the observable.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(N, 3)
ndarrayoffloat
- class espressomd.observables.ParticlePositions(**kwargs)[source]¶
Bases:
ObservableCalculates the particle positions for particles with given ids.
Output format: \((x_1,\ y_1,\ z_1),\ (x_2,\ y_2,\ z_2),\ \dots,\ (x_n,\ y_n,\ z_n)\).
The particles are ordered according to the list of ids passed to the observable.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(N, 3)
ndarrayoffloat
- class espressomd.observables.ParticleVelocities(**kwargs)[source]¶
Bases:
ObservableCalculates the particle velocities for particles with given ids.
Output format: \((v^x_1,\ v^y_1,\ v^z_1),\ (v^x_2,\ v^y_2,\ v^z_2),\ \dots,\ (v^x_n,\ v^y_n,\ v^z_n)\).
The particles are ordered according to the list of ids passed to the observable.
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(N, 3)
ndarrayoffloat
- class espressomd.observables.Pressure(**kwargs)[source]¶
Bases:
ObservableCalculates the total scalar pressure.
- calculate()¶
Run the observable.
- Return type:
float
- class espressomd.observables.PressureTensor(**kwargs)[source]¶
Bases:
ObservableCalculates the total pressure tensor.
- calculate()¶
Run the observable.
- Return type:
(3, 3)
ndarrayoffloat
- class espressomd.observables.ProfileObservable(**kwargs)[source]¶
Bases:
ObservableBase class for histogram-based observables.
- class espressomd.observables.RDF(**kwargs)[source]¶
Bases:
ObservableCalculates a radial distribution function. The result is normalized by the bulk concentration.
- Parameters:
ids1 (array_like of
int) – The ids of (existing) particles to calculate the distance from.ids2 (array_like of
int, optional) – The ids of (existing) particles to calculate the distance to. If not provided, useids1.n_r_bins (
int) – Number of bins in radial direction.min_r (
float) – Minimumrto consider.max_r (
float) – Maximumrto consider.
- calculate()¶
Run the observable.
- Returns:
The RDF.
- Return type:
(
n_r_bins,)ndarrayoffloat
- class espressomd.observables.TotalForce(**kwargs)[source]¶
Bases:
ObservableCalculates the total force on particles with given ids.
Note that virtual sites are not included since forces on them do not enter the equation of motion directly.
Output format: \(\left(\sum_i f^x_i, \sum_i f^y_i, \sum_i f^z_i\right)\)
- Parameters:
ids (array_like of
int) – The ids of (existing) particles to take into account.
- calculate()¶
Run the observable.
- Return type:
(3,)
ndarrayoffloat
espressomd.pair_criteria module¶
- class espressomd.pair_criteria.BondCriterion(**kwargs)[source]¶
Bases:
_PairCriterionPair criterion returning true, if a pair bond of given type exists between them
The following parameters can be passed to the constructor, changed via
set_params()and retrieved viaget_params().- Parameters:
bond_type (
int) – numeric type of the bond
- class espressomd.pair_criteria.DistanceCriterion(**kwargs)[source]¶
Bases:
_PairCriterionPair criterion returning true, if particles are closer than a cutoff. Periodic boundaries are treated via minimum image convention.
The following parameters can be passed to the constructor, changed via
set_params()and retrieved viaget_params().- Parameters:
cut_off (
float) – distance cutoff for the criterion
- class espressomd.pair_criteria.EnergyCriterion(**kwargs)[source]¶
Bases:
_PairCriterionPair criterion returning true, if the short range energy between the particles is superior or equal to the cutoff.
Be aware that the short range energy contains the short range part of dipolar and electrostatic interactions, but not the long range part.
The following parameters can be passed to the constructor, changed via
set_params()and retrieved viaget_params().- Parameters:
cut_off (
float) – energy cutoff for the criterion
espressomd.particle_data module¶
- class espressomd.particle_data.ParticleHandle(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper- id¶
Particle identifier.
- Type:
int
- type¶
The particle type for non-bonded interactions.
Note
The value of
typehas to be an integer >= 0.- Type:
int
- mol_id¶
The molecule id of the particle.
The particle
mol_idis used to differentiate between particles belonging to different molecules, e.g. when virtual sites are used, or object-in-fluid cells. The defaultmol_idfor all particles is 0.Note
The value of
mol_idhas to be an integer >= 0.- Type:
int
- pos¶
The unwrapped (not folded into central box) particle position.
- Type:
(3,) array_like of
float
- pos_folded¶
The wrapped (folded into central box) position vector of a particle.
Note
Setting the folded position is ambiguous and is thus not possible, please use
posinstead.Examples
>>> import espressomd >>> system = espressomd.System(box_l=[10, 10, 10]) >>> system.part.add(pos=(5, 0, 0)) >>> system.part.add(pos=(10, 0, 0)) >>> system.part.add(pos=(25, 0, 0)) >>> for p in system.part: ... print(p.pos) [ 5. 0. 0.] [ 10. 0. 0.] [ 25. 0. 0.] >>> for p in system.part: ... print(p.pos_folded) [5.0, 0.0, 0.0] [0.0, 0.0, 0.0] [5.0, 0.0, 0.0]
- Type:
(3,) array_like of
float
- image_box¶
The image box the particles is in.
This is the number of times the particle position has been folded by the box length in each direction.
- Type:
(3,) array_like of
int
- lees_edwards_offset¶
The accumulated Lees-Edwards offset. Can be used to reconstruct continuous trajectories.
- Type:
float
- lees_edwards_flag¶
The Lees-Edwards flag that indicate if the particle crossed the upper or lower boundary.
- Type:
int
- v¶
The particle velocity in the lab frame.
Note
The velocity will be updated during integration.
- Type:
(3,) array_like of
float
- f¶
The instantaneous force acting on this particle.
Note
The force is recomputed during the integration step and any force set in this way is immediately lost at the next integration step.
- Type:
(3,) array_like of
float
- node¶
The node the particle is on, identified by its MPI rank.
- Type:
(3,) array_like of
int
- mass¶
Particle mass.
- Type:
float
- omega_lab¶
The particle angular velocity the lab frame.
Note
This needs the feature
ROTATION.If you set the angular velocity of the particle in the lab frame, the orientation of the particle (
quat) must be set before settingomega_lab, otherwise the conversion from lab to body frame will not be handled properly.See also
- Type:
(3,) array_like of
float
- quat¶
Quaternion representation of the particle rotational position.
Note
This needs the feature
ROTATION.- Type:
(4,) array_like of
float
- director¶
The particle director.
The
directordefines the the z-axis in the body-fixed frame. If particle rotations happen, the director, i.e., the body-fixed coordinate system co-rotates. Properties such as the angular velocityespressomd.particle_data.ParticleHandle.omega_bodyare evaluated in this body-fixed coordinate system. When using particle dipoles, the dipole moment is co-aligned with the particle director. Setting the director thus modifies the dipole moment orientation (espressomd.particle_data.ParticleHandle.dip) and vice versa. See also Rotational degrees of freedom and particle anisotropy.Note
This needs the feature
ROTATION.- Type:
(3,) array_like of
float
- omega_body¶
The particle angular velocity in body frame.
This property sets the angular momentum of this particle in the particles co-rotating frame (or body frame).
Note
This needs the feature
ROTATION.- Type:
(3,) array_like of
float
- torque_lab¶
The particle torque in the lab frame.
This property defines the torque of this particle in the fixed frame (or laboratory frame).
Note
The orientation of the particle (
quat) must be set before setting this property, otherwise the conversion from lab to body frame will not be handled properly.- Type:
(3,) array_like of
float
- rinertia¶
The particle rotational inertia.
Sets the diagonal elements of this particle’s rotational inertia tensor. These correspond with the inertial moments along the coordinate axes in the particle’s co-rotating coordinate system. When the particle’s quaternions are set to
[1, 0, 0, 0,], the co-rotating and the fixed (lab) frames are co-aligned.Note
This needs the feature
ROTATIONAL_INERTIA.- Type:
(3,) array_like of
float
- q¶
Particle charge.
Note
This needs the feature
ELECTROSTATICS.- Type:
float
- mu_E¶
Particle electrophoretic velocity.
This effectively acts as a velocity offset between a lattice-Boltzmann fluid and the particle. Has only an effect if LB is turned on.
Note
This needs the feature
LB_ELECTROHYDRODYNAMICS.- Type:
float
- virtual¶
Virtual flag.
Declares the particles as virtual (
True) or non-virtual (False, default).Note
This needs the feature
VIRTUAL_SITES- Type:
bool
- vs_quat¶
Virtual site quaternion.
This quaternion describes the virtual particles orientation in the body fixed frame of the related real particle.
Note
This needs the feature
VIRTUAL_SITES_RELATIVE.- Type:
(4,) array_like of
float
- vs_relative¶
Virtual sites relative parameters.
Allows for manual access to the attributes of virtual sites in the “relative” implementation. Format:
(PID, distance, quaternion). PID denotes the id of the particle to which this virtual site is related and distance the distance between non-virtual and virtual particle. The relative orientation is specified as a quaternion.Note
This needs the feature
VIRTUAL_SITES_RELATIVE- Type:
tuple
- dip¶
The orientation of the dipole axis.
Note
This needs the feature
DIPOLES.- Type:
(3,) array_like of
float
- dipm¶
The magnitude of the dipole moment.
Note
This needs the feature
DIPOLES.- Type:
float
- dip_fld¶
Total dipole field value at the position of the particle.
Note
This needs the feature
DIPOLE_FIELD_TRACKING.- Type:
(3,) array_like of
float
- ext_force¶
An additional external force applied to the particle.
Note
This needs the feature
EXTERNAL_FORCES.- Type:
(3,) array_like of
float
- fix¶
Fixes the particle motion in the specified cartesian directions.
Fixes the particle in space. It is possible to fix motion in the x-, y-, or z-direction independently. For example:
part.by_id(1).fix = [False, False, True]
will fix motion for particle with index 1 only in the z-direction.
Note
This needs the feature
EXTERNAL_FORCES.- Type:
(3,) array_like of
bool
- ext_torque¶
An additional external torque is applied to the particle.
Note
This torque is specified in the laboratory frame!
This needs features
EXTERNAL_FORCESandROTATION.
- Type:
(3,) array_like of
float
- gamma¶
The translational frictional coefficient used in the Langevin and Brownian thermostats.
Note
This needs feature
THERMOSTAT_PER_PARTICLEand optionallyPARTICLE_ANISOTROPY.See also
espressomd.thermostat.Thermostat.set_langevin()Setting the parameters of the Langevin thermostat
- Type:
floator (3,) array_like offloat
- gamma_rot¶
The particle rotational frictional coefficient used in the Langevin and Brownian thermostats.
gamma_rot :
floator (3,) array_like offloatNote
This needs features
THERMOSTAT_PER_PARTICLE,ROTATIONand optionallyPARTICLE_ANISOTROPY.- Type:
floator (3,) array_like offloat
- rotation¶
Switches the particle’s rotational degrees of freedom in the Cartesian axes in the body-fixed frame. The content of the torque and omega variables are meaningless for the co-ordinates for which rotation is disabled.
The default is not to integrate any rotational degrees of freedom.
rotation : (3,) array_like of
boolNote
This needs the feature
ROTATION.- Type:
(3,) array_like of
bool
- swimming¶
Set swimming parameters.
This property takes a dictionary with a different number of entries depending whether there is an implicit fluid (i.e. with the Langevin thermostat) of an explicit fluid (with lattice-Boltzmann).
Swimming enables particle self-propulsion in the direction determined by its quaternion. For setting the quaternion of the particle see
quat. Self-propulsion is achieved by imposing a constant force termf_swimalong the particle direction. The steady-state propulsion speed (v_swim) can be calculated from the friction (gamma) of a thermostat:v_swim = f_swim / gamma. When resolving hydrodynamics via lattice-Boltzmann, the swimming attribute can be used to create the typical dipolar flowfield of self-propelled particles: settingis_engine_force_on_fluidtoTruewill make the particle not experience any friction or noise, but instead apply the swim forcef_swimto the fluid. Useespressomd.swimmer_helpers.add_dipole_particle()to automate such a setup.- Parameters:
f_swim (
float) – Magnitude of the self-propulsion force.is_engine_force_on_fluid (
bool) – Default:False. IfTrue, the particle will apply the swimming force to the fluid instead of experiencing drag.
Notes
This needs feature
ENGINE, and optionallyVIRTUAL_SITES_RELATIVEto add the propulsion force on a lattice-Boltzmann fluid.Examples
>>> import espressomd >>> # swimming withut hydrodynamics >>> system = espressomd.System(box_l=[10, 10, 10]) >>> partcl = system.part.add(pos=[1, 0, 0], swimming={'f_swim': 0.03}) >>> # swimming with hydrodynamics >>> import espressomd.swimmer_helpers.add_dipole_particle as add_dip >>> dipole_partcl = add_dip(system, partcl, 2., 0)
- delete_all_bonds()¶
Delete all bonds from the particle.
See also
delete_bondDelete an unverified bond held by the particle.
bondsParticleproperty containing a list of all current bonds held byParticle.
- is_virtual()¶
Whether the particle is a virtual site.
- add_bond(bond)[source]¶
Add a single bond to the particle.
- Parameters:
bond (
tuple) – tuple where the first element is either a bond ID or a bond object, and the next elements are particle ids or particle objects to be bonded to.
See also
bondsParticleproperty containing a list of all current bonds held byParticle.
Examples
>>> import espressomd.interactions >>> >>> system = espressomd.System(box_l=3 * [10]) >>> >>> # define a harmonic potential and add it to the system >>> harm_bond = espressomd.interactions.HarmonicBond(r_0=1, k=5) >>> system.bonded_inter.add(harm_bond) >>> >>> # add two particles >>> p1 = system.part.add(pos=(1, 0, 0)) >>> p2 = system.part.add(pos=(2, 0, 0)) >>> >>> # bond them via the bond type >>> p1.add_bond((harm_bond, p2)) >>> # or via the bond index (zero in this case since it is the first one added) >>> p1.add_bond((0, p2))
- add_exclusion(partner)[source]¶
Exclude non-bonded interactions with the given partner.
Note
This needs the feature
EXCLUSIONS.- Parameters:
partner (
ParticleHandleorint) – Particle to exclude.
- property bonds¶
The bonds stored by this particle. Note that bonds are only stored by one partner. You need to define a bonded interaction.
A bond tuple is specified as a bond identifier associated with a particle
(bond_ID, (*part_ID,)). A single particle may contain multiple bonds.Type: Ragged array.
Note
Bond ids have to be an integer >= 0.
See also
espressomd.particle_data.ParticleHandle.add_bondMethod to add bonds to a
Particleespressomd.particle_data.ParticleHandle.delete_bondMethod to remove bonds from a
Particle
- convert_vector_body_to_space(vec)[source]¶
Convert the given vector from the particle’s body frame to the space frame.
- convert_vector_space_to_body(vec)[source]¶
Convert the given vector from the space frame to the particle’s body frame.
- delete_bond(bond)[source]¶
Delete a single bond from the particle.
- Parameters:
bond (
tuple) – tuple where the first element is either a bond ID or a bond object, and the next elements are particle ids or particle objects that are bonded to.
See also
bondsParticleproperty containing a list of all bonds currently held byParticle.
Examples
>>> import espressomd.interactions >>> >>> system = espressomd.System(box_l=3 * [10]) >>> >>> # define a harmonic potential and add it to the system >>> harm_bond = espressomd.interactions.HarmonicBond(r_0=1, k=5) >>> system.bonded_inter.add(harm_bond) >>> >>> # bond two particles to the first one >>> p0 = system.part.add(pos=(1, 0, 0)) >>> p1 = system.part.add(pos=(2, 0, 0)) >>> p2 = system.part.add(pos=(1, 1, 0)) >>> p0.add_bond((harm_bond, p1)) >>> p0.add_bond((harm_bond, p2)) >>> >>> print(p0.bonds) ((HarmonicBond(0): {'r_0': 1.0, 'k': 5.0, 'r_cut': 0.0}, 1), (HarmonicBond(0): {'r_0': 1.0, 'k': 5.0, 'r_cut': 0.0}, 2)) >>> # delete the first bond >>> p0.delete_bond(p0.bonds[0]) >>> print(p0.bonds) ((HarmonicBond(0): {'r_0': 1.0, 'k': 5.0, 'r_cut': 0.0}, 2),)
- delete_exclusion(partner)[source]¶
Remove exclusion of non-bonded interactions with the given partner.
Note
This needs the feature
EXCLUSIONS.- Parameters:
partner (
ParticleHandleorint) – Particle to remove from exclusions.
- delete_verified_bond(bond)[source]¶
Delete a single bond from the particle. The validity of which has already been verified.
- Parameters:
bond (
tuple) – tuple where the first element is either a bond ID of a bond type, and the last element is the ID of the partner particle to be bonded to.
See also
delete_bondDelete an unverified bond held by the
Particle.bondsParticleproperty containing a list of all current bonds held byParticle.
- property exclusions¶
The exclusion list of particles where non-bonded interactions are ignored.
Note
This needs the feature
EXCLUSIONS.Type: (N,) array_like of
int
- normalize_and_check_bond_or_throw_exception(bond)[source]¶
Checks the validity of the given bond:
If the bondtype is given as an object or a numerical id
If all partners are of type
intIf the number of partners satisfies the bond
If the bond type used exists (is lower than
n_bonded_ia)If the number of bond partners fits the bond type
Throws an exception if any of these are not met.
Normalize the bond, i.e. replace bond ids by bond objects and particle objects by particle ids.
- property propagation¶
- rotate(axis, angle)[source]¶
Rotate the particle around the given axis.
- Parameters:
axis ((3,) array_like of
float)angle (
float)
- to_dict()[source]¶
Returns the particle’s attributes as a dictionary.
It includes the content of
particle_attributes, minus a few exceptions:
- update(new_properties)[source]¶
Update properties of a particle.
- Parameters:
new_properties (
dict) – Map particle property names to values. All properties except for the particle id can be changed.
Examples
>>> import espressomd >>> system = espressomd.System(box_l=[10, 10, 10]) >>> p = system.part.add(pos=[1, 2, 3], q=1, virtual=True) >>> print(p.pos, p.q, p.virtual) [1. 2. 3.] 1.0 True >>> p.update({'pos': [4, 5, 6], 'virtual': False, 'q': 0}) >>> print(p.pos, p.q, p.virtual) [4. 5. 6.] 0.0 False
- vs_auto_relate_to(rel_to, override_cutoff_check=False, couple_to_lb=False, couple_to_langevin=False)[source]¶
Setup this particle as virtual site relative to the particle in argument
rel_to. A particle cannot relate to itself.- Parameters:
rel_to (
intorParticleHandle) – Particle to relate to (either particle id or particle object).override_cutoff_check (
bool) – If True, does not check whether the cell system cutoffs are consistent with the distance between virtual and non-virtual particles.couple_to_lb (
bool) – If True, the virtual site is coupled to LB friction and noise.couple_to_langevin (
bool) – If True, the virtual site is coupled to Langevin friction and noise. Ifcouple_to_lbis also True, propagate LB’s equations of motion and Langevin’s equations of rotation.
- class espressomd.particle_data.ParticleList(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperProvides access to the particles.
- clear()¶
Remove all particles.
- auto_exclusions()¶
Add exclusions between particles that are connected by pair bonds, including virtual bonds. Angle and dihedral bonds are ignored. The most common use case for this method is to auto-exclude virtual sites.
Another use case is to exclude 1-2, 1-3 and optionally 1-4 non-nonded interactions on polymer chains. This technique is commonly used in atomistic molecular dynamics engines such as NAMD, AMBER or GROMACS, where the short-range part of the potential energy surface is better approximated with Fourier sums (using dihedral bonds) than with pair potentials. Linear, branched and circular topologies are supported.
Requires feature
EXCLUSIONS.- Parameters:
distance (
int) – Maximal length of a chain in unit of bonds. The topology will be traversed recursively until the bond chain either terminates or reaches that distance.
- add(*args, **kwargs)[source]¶
Adds one or several particles to the system
- Parameters:
Either a dictionary or a bunch of keyword args.
- Return type:
Returns an instance of
espressomd.particle_data.ParticleHandlefor each added particle.
Examples
>>> import espressomd >>> system = espressomd.System(box_l=[10, 10, 10]) >>> # add two particles >>> system.part.add(id=0, pos=(1, 0, 0)) >>> system.part.add(id=1, pos=(2, 0, 0))
posis mandatory,idcan be omitted, in which case it is assigned automatically. Several particles can be added by passing one value per particle to each property:system.part.add(pos=((1, 2, 3), (4, 5, 6)), q=(1, -1))
- property highest_particle_id¶
Largest particle id.
- select(*args, **kwargs)[source]¶
Generate a particle slice by filtering particles via a user-defined criterion.
Parameters:
Either: a keyword arguments in which the keys are names of particle properties and the values are the values to filter for. E.g.,:
system.part.select(type=0, q=1)
Or: a function taking a ParticleHandle as argument and returning True if the particle is to be filtered for. E.g.,:
system.part.select(lambda p: p.pos[0] < 0.5)
- Returns:
An instance of
ParticleSlicecontaining the selected particles- Return type:
- writevtk(fname, types='all')[source]¶
Write the positions and velocities of particles with specified types to a VTK file.
- Parameters:
fname (
str) – Filename of the target output filetypes (list of
intor the string ‘all’, optional (default: ‘all’)) – A list of particle types which should be output to ‘fname’
Examples
>>> import espressomd >>> system = espressomd.System(box_l=[10, 10, 10]) >>> # add several particles >>> system.part.add(pos=0.5 * system.box_l, v=[1, 0, 0], type=0) >>> system.part.add(pos=0.4 * system.box_l, v=[0, 2, 0], type=1) >>> system.part.add(pos=0.7 * system.box_l, v=[2, 0, 1], type=1) >>> system.part.add(pos=0.1 * system.box_l, v=[0, 0, 1], type=2) >>> # write to VTK >>> system.part.writevtk("part_type_0_1.vtk", types=[0, 1]) >>> system.part.writevtk("part_type_2.vtk", types=[2]) >>> system.part.writevtk("part_all.vtk")
Todo
move to
./io/writer/
- class espressomd.particle_data.ParticleSlice(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperHandle slice inputs. Set values for selected slices or return values as a single list.
- property bonds¶
- property dip¶
- property dip_fld¶
- property dipm¶
- property director¶
- property exclusions¶
- property ext_force¶
- property ext_torque¶
- property f¶
- property fix¶
- property gamma¶
- property gamma_rot¶
- property id¶
- property image_box¶
- property lees_edwards_flag¶
- property lees_edwards_offset¶
- property mass¶
- property mol_id¶
- property mu_E¶
- property node¶
- property omega_body¶
- property omega_lab¶
- property pos¶
- property pos_folded¶
Particle position (folded into central image).
- property propagation¶
- property q¶
- property quat¶
- property rinertia¶
- property rotation¶
- property swimming¶
- to_dict()[source]¶
Returns the particles attributes as a dictionary.
It can be used to save the particle data and recover it by using
>>> p = system.part.add(...) >>> particle_dict = p.to_dict() >>> system.part.add(particle_dict)
It includes the content of
particle_attributes, minus a few exceptions:
- property torque_lab¶
- property type¶
- property v¶
- property vs_quat¶
- property vs_relative¶
espressomd.polymer module¶
- espressomd.polymer.linear_polymer_positions(start_positions=(), min_distance=0.0, respect_constraints=False, max_tries=1000, **kwargs)[source]¶
Generate particle positions for polymer creation.
- Parameters:
n_polymers (
int, required) – Number of polymer chainsbeads_per_chain (
int, required) – Number of monomers per chainbond_length (
float, required) – distance between adjacent monomers in a chainseed (
int, required) – Seed for the RNG used to generate the particle positions.bond_angle (
float, optional) – Angle in radians between adjacent bonds within a polymer. Must be in the range \([0, \pi]\). If omitted, random angles are drawn.start_positions ((N, 3) array_like
float, optional) – If set, this vector defines the start positions for the polymers, i.e., the position of each polymer’s first monomer bead. Here, a numpy array of shape(n_polymers, 3)is expected.min_distance (
float, optional) – Minimum distance between all generated positions.respect_constraints (
bool, optional) – If True, the particle setup tries to obey previously defined constraints.max_tries (
int, optional) – Maximal number of attempts to generate every monomer position, as well as maximal number of retries per polymer, if choosing suitable monomer positions fails. Depending on the total number of beads and constraints, this value needs to be adapted.
- Returns:
Three-dimensional numpy array, namely a list of polymers containing the coordinates of the respective monomers.
- Return type:
ndarray
- espressomd.polymer.setup_diamond_polymer(system=None, bond=None, MPC=0, dist_cM=1, val_cM=0.0, val_nodes=0.0, start_id='auto', no_bonds=False, type_nodes=0, type_nM=1, type_cM=2)[source]¶
Places particles to form a diamond lattice shaped polymer. Can also assign charges and bonds at the appropriate places.
- Parameters:
system (
espressomd.system.System, required) – System to which the particles will be added.bond (
espressomd.interactions.BondedInteraction, required ifno_bonds == False) – The bond to be created between monomers. Should be compatible with the spacingsystem.box_l[0]*(0.25 * sqrt(3))/(MPC + 1)between monomers.no_bonds (
bool, optional) – If True, the particles will only be placed in the system but not connected by bonds. In that case, thebondargument can be omitted. Defaults toFalse.MPC (
int, optional) – Monomers per chain, where chain refers to the connection between the 8 lattice nodes of the diamond lattice. Defaults to 0.dist_cM (
int, optional) – Distance between charged monomers in the chains. Defaults to 1.val_cM (
float, optional) – Valence of the charged monomers in the chains. Defaults to 0.val_nodes (
float, optional) – Valence of the node particles. Defaults to 0.start_id (
intor'auto', optional) – Start id for particle creation. Subsequent ids will be contiguous integers. If'auto', particle ids will start after the highest id of particles already in the system.type_nodes (
int, optional) – Type assigned to the node particles. Defaults to 0.type_nM (
int, optional) – Type assigned to the neutral monomers in the chains. Defaults to 1.type_cM (
int, optional) – Type assigned to the charged monomers in the chains. Defaults to 2.
espressomd.profiler module¶
- class espressomd.profiler.Caliper(*args, **kwargs)[source]¶
Bases:
ScriptInterfaceHelperAdd Caliper section markers at runtime.
- begin_section()¶
Start named section.
- Parameters:
label (
str) – Name of the section.
- end_section()¶
End named section.
- Parameters:
label (
str) – Name of the section.
espressomd.propagation module¶
- class espressomd.propagation.Propagation(value, names=<not given>, *values, module=None, qualname=None, type=None, start=1, boundary=None)[source]¶
Bases:
IntFlagFlags for propagation modes. Use the pipe operator to combine multiple propagation modes. Not all combinations of propagation modes are allowed. Flags for virtual sites are special and instruct the propagator to skip integration; virtual sites can still be coupled to thermostats (Langevin, lattice-Boltzmann) to apply friction and noise to their forces and torques.
- NONE = 0¶
No propagation.
- SYSTEM_DEFAULT = 1¶
Use the system default propagator for motion and rotation.
- TRANS_NEWTON = 2¶
Velocity-Verlet algorithm that integrates Newton’s equations of motion.
- TRANS_LANGEVIN = 4¶
Velocity-Verlet algorithm that integrates Langevin’s equations of motion.
- TRANS_LANGEVIN_NPT = 8¶
Velocity-Verlet algorithm that integrates Langevin’s equations of motion coupled to a piston.
- TRANS_VS_RELATIVE = 16¶
Algorithm for virtual sites relative motion.
- TRANS_LB_MOMENTUM_EXCHANGE = 32¶
Algorithm for momentum exchange between particles and LB fluid cells.
- TRANS_LB_TRACER = 64¶
Algorithm for LB inertialess tracers advection.
- TRANS_BROWNIAN = 128¶
Euler algorithm that integrates Brownian’s equations of motion.
- TRANS_STOKESIAN = 256¶
Euler algorithm that integrates Stoke’s equations of motion.
- ROT_EULER = 1024¶
Velocity-Verlet algorithm that integrates Euler’s equations of rotation.
- ROT_LANGEVIN = 2048¶
Velocity-Verlet algorithm that integrates Langevin’s equations of rotation.
- ROT_VS_RELATIVE = 4096¶
Algorithm for virtual sites relative rotation.
- ROT_BROWNIAN = 8192¶
Euler algorithm that integrates Brownian’s equations of rotation.
- ROT_STOKESIAN = 16384¶
Euler algorithm that integrates Stokes’ equations of rotation.
espressomd.reaction_methods module¶
- class espressomd.reaction_methods.ConstantpHEnsemble(**kwargs)[source]¶
Bases:
ReactionAlgorithmThis class implements the constant pH Ensemble.
When adding an acid-base reaction, the acid and base particle types are always assumed to be at index 0 of the lists passed to arguments
reactant_typesandproduct_types.- constant_pH¶
Constant pH value.
- Type:
float
- add_reaction(*args, **kwargs)[source]¶
Sets up a reaction in the forward and backward direction.
- Parameters:
gamma (
float) – Equilibrium constant \(\Gamma\) of the reaction in simulation units (see section Reaction ensemble for its definition).reactant_types (list of
int) – List of particle types of reactants in the reaction.reactant_coefficients (list of
int) – List of stoichiometric coefficients of the reactants in the same order as the list of their types.product_types (list of
int) – List of particle types of products in the reaction.product_coefficients (list of
int) – List of stoichiometric coefficients of products of the reaction in the same order as the list of their typesdefault_charges (
dict) – A dictionary of default charges for types that occur in the provided reaction.check_for_electroneutrality (
bool) – Check for electroneutrality of the given reaction. Default isTrue.
- calculate_acceptance_probability(reaction_id, E_pot_diff)[source]¶
Calculate the acceptance probability of a Monte Carlo move.
- Parameters:
reaction_id (
int) – Identifier of the reaction that was carried out in the move.E_pot_diff (
float) – The potential energy difference for the move.
- Returns:
The acceptance probability.
- Return type:
float
- class espressomd.reaction_methods.ReactionAlgorithm(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperThis class provides the base class for Reaction Algorithms like the Reaction Ensemble algorithm and the constant pH method. Initialize the reaction algorithm by setting the standard pressure, temperature, and the exclusion range.
Note: When creating particles the velocities of the new particles are set according the Maxwell-Boltzmann distribution. In this step the mass of the new particle is assumed to equal 1.
- Parameters:
kT (
float) – Thermal energy of the system in simulation unitsexclusion_range (
float) – Minimal distance from any particle, within which new particles will not be inserted.seed (
int) – Initial counter value (or seed) of the Mersenne Twister RNG.exclusion_radius_per_type (
dict, optional) – Mapping of particle types to exclusion radii.search_algorithm (
str) – Pair search algorithm. Default is"order_n", which evaluates the distance between the inserted particle and all other particles in the system, which scales with O(N). For MPI-parallel simulations, the"parallel"method is faster. The"parallel"method is not recommended for simulations on 1 MPI rank, since it comes with the overhead of a ghost particle update.
- remove_constraint()¶
Remove any previously defined constraint. Requires setting the volume using
set_volume().
- set_cylindrical_constraint_in_z_direction()¶
Constrain the reaction moves within a cylinder aligned on the z-axis. Requires setting the volume using
set_volume().- Parameters:
center_x (
float) – x coordinate of center of the cylinder.center_y (
float) – y coordinate of center of the cylinder.radius_of_cylinder (
float) – radius of the cylinder
- set_wall_constraints_in_z_direction()¶
Restrict the sampling area to a slab in z-direction. Requires setting the volume using
set_volume(). This constraint is necessary when working with Electrostatic Layer Correction (ELC).- Parameters:
slab_start_z (
float) – z coordinate of the bottom wall.slab_end_z (
float) – z coordinate of the top wall.
Examples
>>> import espressomd >>> import espressomd.shapes >>> import espressomd.electrostatics >>> import espressomd.reaction_methods >>> import numpy as np >>> # setup a charged system >>> box_l = 20 >>> elc_gap = 10 >>> system = espressomd.System(box_l=[box_l, box_l, box_l + elc_gap]) >>> system.time_step = 0.001 >>> system.cell_system.skin = 0.4 >>> types = {"HA": 0, "A-": 1, "H+": 2, "wall": 3} >>> charges = {types["HA"]: 0, types["A-"]: -1, types["H+"]: +1} >>> for i in range(10): ... system.part.add(pos=np.random.random(3) * box_l, type=types["A-"], q=charges[types["A-"]]) ... system.part.add(pos=np.random.random(3) * box_l, type=types["H+"], q=charges[types["H+"]]) >>> for particle_type in charges.keys(): ... system.non_bonded_inter[particle_type, types["wall"]].wca.set_params(epsilon=1.0, sigma=1.0) >>> # add ELC actor >>> p3m = espressomd.electrostatics.P3M(prefactor=1.0, accuracy=1e-2) >>> elc = espressomd.electrostatics.ELC(actor=p3m, maxPWerror=1.0, gap_size=elc_gap) >>> system.actors.add(elc) >>> # add constant pH method >>> RE = espressomd.reaction_methods.ConstantpHEnsemble(kT=1, exclusion_range=1, seed=77) >>> RE.constant_pH = 2 >>> RE.add_reaction(gamma=0.0088, reactant_types=[types["HA"]], ... product_types=[types["A-"], types["H+"]], ... default_charges=charges) >>> # add walls for the ELC gap >>> RE.set_wall_constraints_in_z_direction(0, box_l) >>> RE.set_volume(box_l**3) >>> system.constraints.add(shape=espressomd.shapes.Wall(dist=0, normal=[0, 0, 1]), ... particle_type=types["wall"]) >>> system.constraints.add(shape=espressomd.shapes.Wall(dist=-box_l, normal=[0, 0, -1]), ... particle_type=types["wall"])
- get_wall_constraints_in_z_direction()¶
Returns the restrictions of the sampling area in z-direction.
- set_volume()¶
Set the volume to be used in the acceptance probability of the reaction ensemble. This can be useful when using constraints, if the relevant volume is different from the box volume. If not used the default volume which is used, is the box volume.
- Parameters:
volume (
float) – Volume of the system in simulation units
- get_volume()¶
Get the volume to be used in the acceptance probability of the reaction ensemble.
- get_acceptance_rate_configurational_moves()¶
Returns the acceptance rate for the configuration moves.
- get_acceptance_rate_reaction()¶
Returns the acceptance rate for the given reaction.
- Parameters:
reaction_id (
int) – Reaction id
- set_non_interacting_type()¶
Sets the particle type for non-interacting particles. Default value: 100. This is used to temporarily hide particles during a reaction trial move, if they are to be deleted after the move is accepted. Please change this value if you intend to use the type 100 for some other particle types with interactions, or if you need improved performance, as the default value of 100 causes some overhead. Please also note that particles in the current implementation of the Reaction Ensemble are only hidden with respect to Lennard-Jones and Coulomb interactions. Hiding of other interactions, for example a magnetic, needs to be implemented in the code.
- Parameters:
type (
int) – Particle type for the hidden particles
- get_non_interacting_type()¶
Returns the type which is used for hiding particle
- displacement_mc_move_for_particles_of_type()¶
Performs displacement Monte Carlo moves for particles of a given type. New positions of the displaced particles are chosen from the whole box with a uniform probability distribution and new velocities are sampled from the Maxwell-Boltzmann distribution.
The sequence of moves is only accepted if each individual move in the sequence was accepted. Particles are sampled without replacement. Therefore, calling this method once for 10 particles is not equivalent to calling this method 10 times for 1 particle.
- Parameters:
type_mc (
int) – Particle type which should be movedparticle_number_to_be_changed (
int) – Number of particles to move, defaults to 1. Particles are selected without replacement.
- Returns:
Whether all moves were accepted.
- Return type:
bool
- delete_particle()¶
Deletes the particle of the given p_id and makes sure that the particle range has no holes. This function has some restrictions, as e.g. bonds are not deleted. Therefore only apply this function to simple ions.
- Parameters:
p_id (
int) – Id of the particle to be deleted.
- change_reaction_constant()¶
Changes the reaction constant of a given reaction (for both the forward and backward reactions). The
reaction_idwhich is assigned to a reaction depends on the order in whichadd_reaction()was called. The 0th reaction hasreaction_id=0, the next added reaction needs to be addressed withreaction_id=1, etc.- Parameters:
reaction_id (
int) – Identifier of the reaction to modify. Will be multiplied by 2 internally!gamma (
float) – New reaction constant for the forward reaction.
- add_reaction(**kwargs)[source]¶
Sets up a reaction in the forward and backward direction.
- Parameters:
gamma (
float) – Equilibrium constant \(\Gamma\) of the reaction in simulation units (see section Reaction ensemble for its definition).reactant_types (list of
int) – List of particle types of reactants in the reaction.reactant_coefficients (list of
int) – List of stoichiometric coefficients of the reactants in the same order as the list of their types.product_types (list of
int) – List of particle types of products in the reaction.product_coefficients (list of
int) – List of stoichiometric coefficients of products of the reaction in the same order as the list of their typesdefault_charges (
dict) – A dictionary of default charges for types that occur in the provided reaction.check_for_electroneutrality (
bool) – Check for electroneutrality of the given reaction. Default isTrue.
- calculate_acceptance_probability(reaction_id, E_pot_diff)[source]¶
Calculate the acceptance probability of a Monte Carlo move.
- Parameters:
reaction_id (
int) – Identifier of the reaction that was carried out in the move.E_pot_diff (
float) – The potential energy difference for the move.
- Returns:
The acceptance probability.
- Return type:
float
- delete_reaction(**kwargs)[source]¶
Delete a reaction from the set of used reactions (the forward and backward reaction). The
reaction_idwhich is assigned to a reaction depends on the order in whichadd_reaction()was called. The 0th reaction hasreaction_id=0, the next added reaction needs to be addressed withreaction_id=1, etc. After the deletion of a reaction subsequent reactions take thereaction_idof the deleted reaction.- Parameters:
reaction_id (
int) – Reaction id
- generic_oneway_reaction(reaction_id, E_pot_old)[source]¶
Carry out a generic one-way chemical reaction of the type A+B+…+G +… –> K+…X + Z +…
You need to use 2A –> B instead of A+A –> B since in the latter you assume distinctness of the particles, however both ways to describe the reaction are equivalent in the thermodynamic limit (large particle numbers). Furthermore, the order of the reactant and product types matters since particles will be replaced in that order! If there are less reactants than products, new product particles are created randomly in the box. Reactants get their type and charge changed to the corresponding type and charge of the products. If there are more reactants than products, excess reactant particles are deleted.
- Parameters:
reaction_id (
int) – Identifier of the reaction to attempt.E_pot_old (
float) – The current potential energy.
- Returns:
E_pot_new – The potential energy after the move.
- Return type:
float
- get_status()[source]¶
Returns the status of the reaction ensemble in a dictionary containing the used reactions, the used kT and the used exclusion radius.
- class espressomd.reaction_methods.ReactionEnsemble(**kwargs)[source]¶
Bases:
ReactionAlgorithmThis class implements the Reaction Ensemble.
- class espressomd.reaction_methods.SingleReaction(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper
- class espressomd.reaction_methods.WidomInsertion(**kwargs)[source]¶
Bases:
ReactionAlgorithmThis class implements the Widom insertion method in the canonical ensemble for homogeneous systems, where the excess chemical potential is not depending on the location.
- add_reaction(**kwargs)[source]¶
Sets up a reaction in the forward and backward direction.
- Parameters:
gamma (
float) – Equilibrium constant \(\Gamma\) of the reaction in simulation units (see section Reaction ensemble for its definition).reactant_types (list of
int) – List of particle types of reactants in the reaction.reactant_coefficients (list of
int) – List of stoichiometric coefficients of the reactants in the same order as the list of their types.product_types (list of
int) – List of particle types of products in the reaction.product_coefficients (list of
int) – List of stoichiometric coefficients of products of the reaction in the same order as the list of their typesdefault_charges (
dict) – A dictionary of default charges for types that occur in the provided reaction.check_for_electroneutrality (
bool) – Check for electroneutrality of the given reaction. Default isTrue.
- calculate_excess_chemical_potential(**kwargs)[source]¶
Given a set of samples of the particle insertion potential energy, calculates the excess chemical potential and its statistical error.
- Parameters:
particle_insertion_potential_energy_samples (array_like of
float) – Samples of the particle insertion potential energy.N_blocks (
int, optional) – Number of bins for binning analysis.
- Returns:
mean (
float) – Mean excess chemical potential.error (
float) – Standard error of the mean.
- calculate_particle_insertion_potential_energy(**kwargs)[source]¶
Measures the potential energy when particles are inserted in the system following the reaction provided in
reaction_id. Please define the insertion moves by calling the methodadd_reaction()(with only product types specified).Note that although this function does not provide directly the chemical potential, it can be used to calculate it. For an example of such an application please check
/samples/widom_insertion.py.- Parameters:
reaction_id (
int) – Reaction identifier. Will be multiplied by 2 internally to skip reverse reactions, i.e. deletion reactions!- Returns:
The particle insertion potential energy.
- Return type:
float
espressomd.rotation module¶
- espressomd.rotation.diagonalized_inertia_tensor(positions, masses)[source]¶
Calculate the diagonalized inertia tensor with respect to the center of mass for given point masses at given positions.
- Parameters:
positions ((N,3) array_like of
float) – Positions of the masses.masses ((N,) array_like of
float)
- Returns:
(3,) array_like of
float– Principal moments of inertia.(3,3) array_like of
float– Principal axes of inertia. Note that the second axis is the coordinate axis (same as input).
- espressomd.rotation.inertia_tensor(positions, masses)[source]¶
Calculate the inertia tensor for given point masses at given positions.
- Parameters:
positions ((N,3) array_like of
float) – Point masses’ positions.masses ((N,) array_like of
float)
- Returns:
Moment of inertia tensor.
- Return type:
(3,3) array_like of
float
Notes
See wikipedia.
- espressomd.rotation.matrix_to_quat(m)[source]¶
Convert the (proper) rotation matrix to the corresponding quaternion representation based on pyquaternion.
- Parameters:
m ((3,3) array_like of
float) – Rotation matrix.- Returns:
Quaternion representation of the rotation.
- Return type:
array_like of
float- Raises:
ValueError – If the matrix does not a correspond to a proper rotation (det(m) != 1).
espressomd.script_interface module¶
- class espressomd.script_interface.PScriptInterface(name=None, policy='GLOBAL', sip=None, **kwargs)¶
Bases:
objectPython interface to a core ScriptInterface object. The core ScriptInterface class is itself an interface for other core classes, such as constraints, shapes, observables, etc.
This class can be instantiated in two ways: (1) with the object id of an existing core ScriptInterface object, or (2) with parameters to construct a new ScriptInterface object in the core.
- Parameters:
sip (
PObjectRef) – Object id of an existing core object (method 1).name (
str) – Name of the core class to instantiate (method 2).**kwargs – Parameters for the core class constructor (method 2).
policy (
str, {‘GLOBAL’, ‘LOCAL’}) – Creation policy. The managed object exists either on all MPI nodes with ‘GLOBAL’ (default), or only on the head node with ‘LOCAL’.
- sip¶
Pointer to a ScriptInterface object in the core.
- Type:
- call_method(self, method, handle_errors_message=None, with_nogil=False, **kwargs)¶
Call a method of the core class.
- Parameters:
method (
str) – Name of the core method.handle_errors_message (
str, optional) – Custom error message for runtime errors raised in a MPI context.with_nogil (
bool, optional) – Run the core method without the GIL ifTrue.**kwargs – Arguments for the method.
- get_parameter(self, name)¶
- get_params(self)¶
- get_sip(self)¶
Get pointer to the core object.
- name(self)¶
Return name of the core class.
- set_params(self, **kwargs)¶
- class espressomd.script_interface.ScriptInterfaceHelper(**kwargs)¶
Bases:
PScriptInterface- define_bound_methods(self)¶
- generate_caller(self, method_name)¶
- class espressomd.script_interface.ScriptObjectList(**kwargs)¶
Bases:
ScriptInterfaceHelperBase class for container-like classes such as
Constraints. Derived classes must implement anadd()method which adds a single item to the container.The core objects must be managed by a container derived from
ScriptInterface::ObjectList.
- class espressomd.script_interface.ScriptObjectMap(**kwargs)¶
Bases:
ScriptInterfaceHelperBase class for container-like classes such as
BondedInteractions. Derived classes must implement anadd()method which adds a single item to the container.The core objects must be managed by a container derived from
ScriptInterface::ObjectMap.- clear(self)¶
Remove all elements.
- items(self)¶
- keys(self)¶
- remove(self, key)¶
Remove the element with the given key. This is a no-op if the key does not exist.
- class espressomd.script_interface.array_variant(input_array)¶
Bases:
ndarrayReturns a numpy.ndarray that will be serialized as a
std::vector.
- espressomd.script_interface.script_interface_register(c)¶
Decorator used to register script interface classes. This will store a name<->class relationship in a registry, so that parameters of type object can be instantiated as the correct python class.
espressomd.shapes module¶
- class espressomd.shapes.Cylinder(**kwargs)[source]¶
Bases:
Shape,ScriptInterfaceHelperA cylinder shape.
- center¶
Coordinates of the center of the cylinder.
- Type:
(3,) array_like of
float
- axis¶
Axis of the cylinder.
- Type:
(3,) array_like of
float
- radius¶
Radius of the cylinder.
- Type:
float
- length¶
Length of the cylinder.
- Type:
float
- direction¶
Surface orientation, for +1 the normal points out of the mantel, for -1 it points inward.
- Type:
int
- open¶
cylinder is open or has caps.
- Type:
bool
- class espressomd.shapes.Ellipsoid(**kwargs)[source]¶
Bases:
Shape,ScriptInterfaceHelperAn ellipsoid.
For now only ellipsoids of revolution are supported. The symmetry axis is aligned parallel to the x-direction.
- center¶
Coordinates of the center of the ellipsoid.
- Type:
(3,) array_like of
float
- a¶
Semiaxis along the axis of rotational symmetry.
- Type:
float
- b¶
Equatorial semiaxes.
- Type:
float
- direction¶
Surface orientation, for +1 the normal points out of the mantel, for -1 it points inward.
- Type:
int
- class espressomd.shapes.HollowConicalFrustum(**kwargs)[source]¶
Bases:
Shape,ScriptInterfaceHelperHollow conical frustum shape.
- cyl_transform_params¶
Parameters of the spacial orientation of the frustum. Contained must be parameters for
centerandaxis.orientationhas no effect, unlesscentral_angle != 0
- r1¶
Radius r1.
- Type:
float
- r2¶
Radius r2.
- Type:
float
- length¶
Length of the conical frustum along
axis.- Type:
float
- thickness¶
The thickness of the frustum. Also determines the rounding radius of the edges
- Type:
float
- direction¶
Surface orientation, for +1 the normal points out of the mantel, for -1 it points inside of the shape. Defaults to 1
- Type:
int, optional
- central_angle¶
A
central_anglecreates an opening in the frustum along the side, centered symmetrically around thedirectionofcyl_transform_params. Must be between0and2 pi. Defaults to 0.- Type:
float, optional
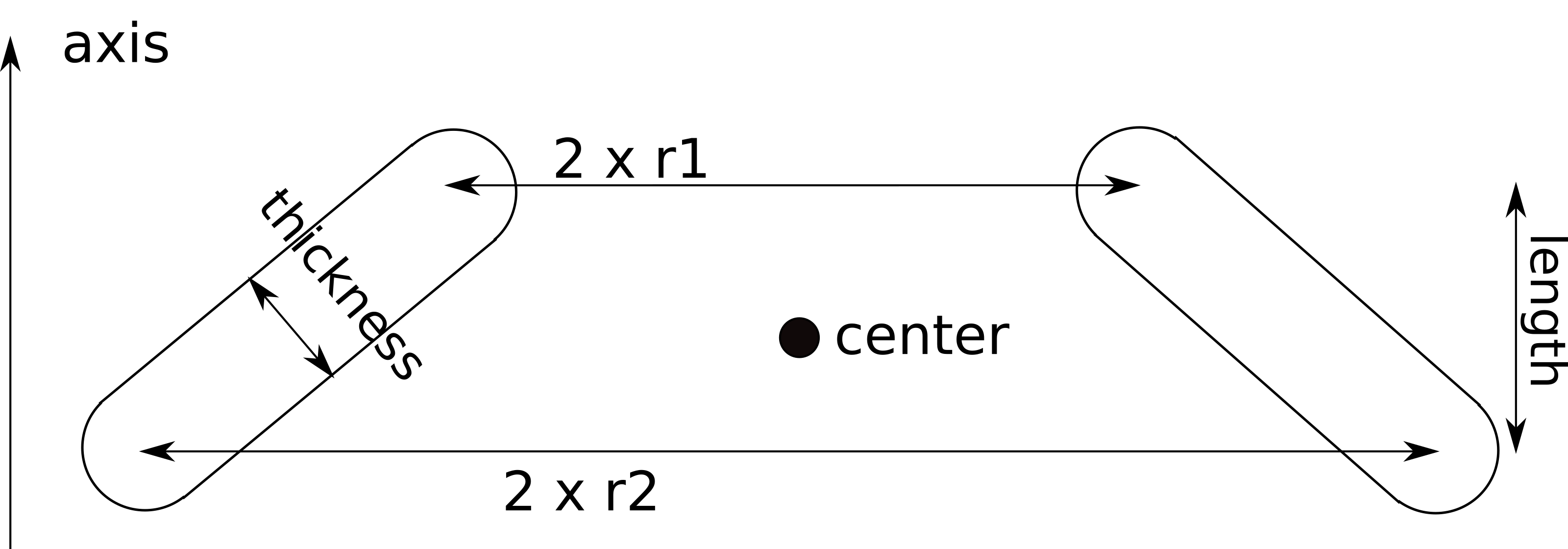
- class espressomd.shapes.Rhomboid(**kwargs)[source]¶
Bases:
Shape,ScriptInterfaceHelperA parallelepiped.
- a¶
First base vector.
- Type:
(3,) array_like of
float
- b¶
Second base vector.
- Type:
(3,) array_like of
float
- c¶
Third base vector.
- Type:
(3,) array_like of
float
- corner¶
Lower left corner of the rhomboid.
- Type:
(3,) array_like of
float
- direction¶
Surface orientation, for +1 the normal points out of the mantel, for -1 it points inward.
- Type:
int
- class espressomd.shapes.SimplePore(**kwargs)[source]¶
Bases:
Shape,ScriptInterfaceHelperTwo parallel infinite planes, and a cylindrical channel connecting them. The cylinder and the planes are connected by torus segments with an adjustable radius.
- radius¶
The radius of the pore.
- Type:
float
- length¶
The distance between the planes.
- Type:
float
- smoothing_radius¶
Radius of the torus segments
- Type:
float
- axis¶
Axis of the cylinder and normal of the planes
- Type:
(3,) array_like of
float
- center¶
Position of the center of the cylinder.
- Type:
(3,) array_like of
float
- class espressomd.shapes.Slitpore(**kwargs)[source]¶
Bases:
Shape,ScriptInterfaceHelper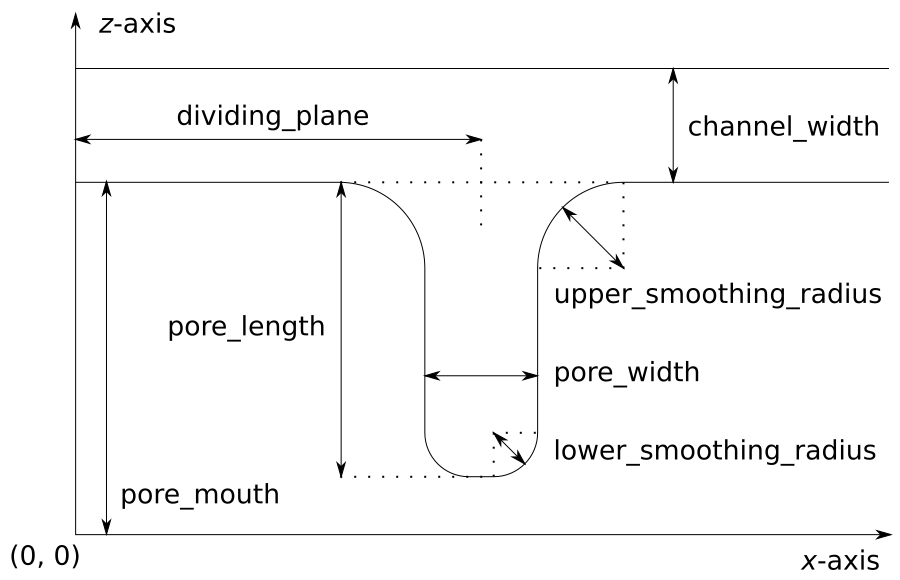
- channel_width¶
- Type:
float
- lower_smoothing_radius¶
- Type:
float
- pore_length¶
- Type:
float
- pore_mouth¶
- Type:
float
- pore_width¶
- Type:
float
- upper_smoothing_radius¶
- Type:
float
- dividing_plane¶
- Type:
float
- class espressomd.shapes.Sphere(**kwargs)[source]¶
Bases:
Shape,ScriptInterfaceHelperA sphere.
- center¶
Center of the sphere
- Type:
(3,) array_like of
float
- radius¶
Radius of the sphere.
- Type:
float
- direction¶
Surface orientation, for +1 the normal points out of the mantel, for -1 it points inward.
- Type:
int
- class espressomd.shapes.SpheroCylinder(**kwargs)[source]¶
Bases:
Shape,ScriptInterfaceHelperA cylinder with hemispheres as caps.
- center¶
Coordinates of the center of the cylinder.
- Type:
(3,) array_like of
float
- axis¶
Axis of the cylinder.
- Type:
(3,) array_like of
float
- radius¶
Radius of the cylinder.
- Type:
float
- direction¶
Surface orientation, for +1 the normal points out of the mantel, for -1 it points inward.
- Type:
int
- length¶
Length of the cylinder (not including the caps).
- Type:
float
- class espressomd.shapes.Torus(**kwargs)[source]¶
Bases:
Shape,ScriptInterfaceHelperA torus shape.
- center¶
Coordinates of the center of the torus.
- Type:
(3,) array_like of
float
- normal¶
Normal axis of the torus.
- Type:
(3,) array_like of
float
- radius¶
Radius of the torus.
- Type:
float
- tube_radius¶
Radius of the tube.
- Type:
float
- direction¶
Surface orientation, for +1 the normal points out of the mantel, for -1 it points inward.
- Type:
int
- class espressomd.shapes.Union(**kwargs)[source]¶
Bases:
Shape,ScriptObjectListA union of shapes.
This shape represents a union of shapes where the distance to the union is defined by the smallest distance to any shape contained in the union.
- size()¶
Number of shapes contained in the union.
- clear()¶
Remove all shapes from the union.
- add(shape)[source]¶
Add a shape to the union.
- Parameters:
shape (array_like / instance of
espressomd.shapes.Shape) – Shape instance(s) to be added to the union.
- remove(shape)[source]¶
Remove a shape from the union.
- Parameters:
shape (array_like / instance of
espressomd.shapes.Shape) – Shape instance(s) to be removed from the union.
espressomd.swimmer_helpers module¶
- espressomd.swimmer_helpers.add_dipole_particle(system, particle, dipole_length, dipole_particle_type, mode='pusher')[source]¶
Add a virtual particle pointing opposite of the swimmer particle either in front of (mode=”puller”) or behind (mode=”pusher”) the swimmer. The virtual particle is set up to exert the swimming force of the swimmer particle on a fluid in the opposite direction.
- Parameters:
system (
System) – The system that the particle belongs to.particle (
ParticleHandle) – The swimmer particle that will be paired with the virtual dipole particle. Must have theswimmingattribute already set up to get the correct value forf_swim.dipole_length (
float) – The distance between the swimmer and the virtual dipole particle.dipole_particle_type (
int) – The type of the virtual dipole particle.mode (
str) – Allowed values: “pusher” (default) and “puller”. Determines whether the virtual dipole particle will be placed in front of or behind the swimmer.
- Returns:
dipole_particle – The newly created particle.
- Return type:
espressomd.system module¶
- class espressomd.system.System(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperThe ESPResSo system class.
- analysis¶
- auto_update_accumulators¶
- bond_breakage¶
- bonded_inter¶
- cell_system¶
- collision_detection¶
- comfixed¶
- constraints¶
- cuda_init_handle¶
- galilei¶
- integrator¶
- lees_edwards¶
- non_bonded_inter¶
- part¶
- thermostat¶
- box_l¶
Dimensions of the simulation box.
- Type:
(3,) array_like of
float
- periodicity¶
System periodicity in
[x, y, z],Falsefor no periodicity in this direction,Truefor periodicity- Type:
(3,) array_like of
bool
- min_global_cut¶
Minimal interaction cutoff.
- Type:
float
- setup_type_map()¶
For using ESPResSo conveniently for simulations in the grand canonical ensemble, or other purposes, when particles of certain types are created and deleted frequently. Particle ids can be stored in lists for each individual type and so random ids of particles of a certain type can be drawn. If you want ESPResSo to keep track of particle ids of a certain type you have to initialize the method by calling the setup function. After that ESPResSo will keep track of particle ids of that type.
- Parameters:
type_list (array_like of
int) – Types to track.
- number_of_particles()¶
Count the number of particles of a given type.
- Parameters:
type (
int(type)) – Particle type to count the number for.- Returns:
The number of particles which have the given type.
- Return type:
int- Raises:
RuntimeError – If the particle
typeis not currently tracked by the system. To select which particle types are tracked, callsetup_type_map().
- rotate_system()¶
Rotate the particles in the system about the center of mass.
If
ROTATIONis activated, the internal rotation degrees of freedom are rotated accordingly.- Parameters:
phi (
float) – Angle between the z-axis and the rotation axis.theta (
float) – Rotation of the axis around the y-axis.alpha (
float) – How much to rotate
- auto_exclusions(distance)[source]¶
Add exclusions between particles that are bonded.
This only considers pair bonds.
Requires feature
EXCLUSIONS.- Parameters:
distance (
int) – Bond distance up to which the exclusions should be added.
- change_volume_and_rescale_particles(d_new, dir='xyz')[source]¶
Change box size and rescale particle coordinates.
- Parameters:
d_new (
float) – New box lengthdir (
str, optional) – Coordinate to work on,"x","y","z"or"xyz"for isotropic. Isotropic assumes a cubic box.
- distance(p1, p2)[source]¶
Return the scalar distance between particles, between a particle and a point or between two points, respecting periodic boundaries.
- Parameters:
p1 (
ParticleHandleor (3,) array_like offloat) – First particle or position.p2 (
ParticleHandleor (3,) array_like offloat) – Second particle or position.
- distance_vec(p1, p2)[source]¶
Return the distance vector between particles, between a particle and a point or between two points, respecting periodic boundaries.
- Parameters:
p1 (
ParticleHandleor (3,) array_like offloat) – First particle or position.p2 (
ParticleHandleor (3,) array_like offloat) – Second particle or position.
- property ekcontainer¶
EK system (diffusion-advection-reaction models).
- property force_cap¶
If > 0, the magnitude of the force on the particles are capped to this value.
Type:
float
- property lb¶
LB solver.
- property max_cut_bonded¶
Maximal cutoff for bonded interactions.
Type:
float
- property max_cut_nonbonded¶
Maximal cutoff for non-bonded interactions.
Type:
float
- property time¶
Total simulation time.
Type:
float
- property time_step¶
MD time step.
Type:
float
- velocity_difference(p1, p2)[source]¶
Return the velocity difference between two particles, considering Lees-Edwards boundary conditions, if active.
- Parameters:
p1 (
ParticleHandle)p2 (
ParticleHandle)
espressomd.thermostat module¶
- class espressomd.thermostat.Brownian(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper
- class espressomd.thermostat.DPDThermostat(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper
- class espressomd.thermostat.IsotropicNpt(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper
- class espressomd.thermostat.LBThermostat(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper
- class espressomd.thermostat.Langevin(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper
- class espressomd.thermostat.Stokesian(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper
- class espressomd.thermostat.ThermalizedBond(**kwargs)[source]¶
Bases:
ScriptInterfaceHelper
- class espressomd.thermostat.Thermostat(**kwargs)[source]¶
Bases:
ScriptInterfaceHelperContainer for the system thermostat. Multiple thermostats can be active simultaneously. When setting up a thermostat raises an exception, e.g. due to invalid parameters, the thermostat gets disabled. Therefore, updating an already active thermostat with invalid parameters will deactivate it.
- turn_off()¶
Turn off all thermostats.
- set_langevin()¶
Set the Langevin thermostat.
- Parameters:
kT (
float) – Thermal energy of the simulated heat bath.gamma (
float) – Contains the friction coefficient of the bath. If the featurePARTICLE_ANISOTROPYis compiled in, thengammacan be a list of three positive floats, for the friction coefficient in each cardinal direction.gamma_rotation (
float, optional) – The same applies togamma_rotation, which requires the featureROTATIONto work properly. But also accepts three floats ifPARTICLE_ANISOTROPYis also compiled in.seed (
int) – Initial counter value (or seed) of the philox RNG. Required on first activation of the Langevin thermostat. Must be positive.
- set_brownian()¶
Set the Brownian thermostat.
- Parameters:
kT (
float) – Thermal energy of the simulated heat bath.gamma (
float) – Contains the friction coefficient of the bath. If the featurePARTICLE_ANISOTROPYis compiled in, thengammacan be a list of three positive floats, for the friction coefficient in each cardinal direction.gamma_rotation (
float, optional) – The same applies togamma_rotation, which requires the featureROTATIONto work properly. But also accepts three floats ifPARTICLE_ANISOTROPYis also compiled in.seed (
int) – Initial counter value (or seed) of the philox RNG. Required on first activation of the Brownian thermostat. Must be positive.
- set_lb()¶
Set the LB thermostat. The kT parameter is automatically extracted from the currently active LB solver.
This thermostat requires the feature
WALBERLA.- Parameters:
seed (
int) – Seed for the random number generator, required if kT > 0. Must be positive.gamma (
float) – Frictional coupling constant for the MD particle coupling.
- set_npt()¶
Set the NPT thermostat.
- Parameters:
kT (
float) – Thermal energy of the heat bath.gamma0 (
float) – Friction coefficient of the bath.gammav (
float) – Artificial friction coefficient for the volume fluctuations.seed (
int) – Initial counter value (or seed) of the philox RNG. Required on first activation of the Langevin thermostat. Must be positive.
- set_dpd()¶
Set the DPD thermostat with required parameters ‘kT’. This also activates the DPD interactions.
- Parameters:
kT (
float) – Thermal energy of the heat bath.seed (
int) – Initial counter value (or seed) of the philox RNG. Required on first activation of the DPD thermostat. Must be positive.
- set_stokesian()¶
Set the SD thermostat with required parameters.
This thermostat requires the feature
STOKESIAN_DYNAMICS.- Parameters:
kT (
float, optional) – Temperature.seed (
int, optional) – Seed for the random number generator.
espressomd.utils module¶
- class espressomd.utils.array_locked(input_array)¶
Bases:
ndarrayReturns a non-writable numpy.ndarray with a special error message upon usage of __setitem__ or in-place operators. Cast return in __get__ of array properties to array_locked to prevent these operations.
- ERR_MSG = 'ESPResSo array properties return non-writable arrays and can only be modified as a whole, not in-place or component-wise. Use numpy.copy(<ESPResSo array property>) to get a writable copy.'¶
- espressomd.utils.check_required_keys(required_keys, obtained_keys)¶
- espressomd.utils.check_type_or_throw_except(x, n, t, msg)¶
Check that
xis of typetand thatnvalues are given, otherwise raise aValueErrorwith messagemsg. Ifxis an array/list/tuple, the type checking is done on the elements, and all elements are checked. Ifnis 1,xis assumed to be a scalar. Integers are accepted when a float was asked for.
- espressomd.utils.check_valid_keys(valid_keys, obtained_keys)¶
- espressomd.utils.handle_errors(msg)¶
Gathers runtime errors.
- Parameters:
msg (
str) – Error message that is to be raised.
- espressomd.utils.is_valid_type(value, t)¶
Extended checks for numpy int, float and bool types. Handles 0-dimensional arrays.
- espressomd.utils.nesting_level(obj)¶
Returns the maximal nesting level of an object.
- espressomd.utils.to_char_pointer(s)¶
Returns a Cython bytes object which contains the information of the provided Python string. Cython bytes objects implicitly cast to raw char pointers.
- Parameters:
s (
str)
- espressomd.utils.to_str(s)¶
Returns a python string.
- Parameters:
s (char*)
espressomd.version module¶
- espressomd.version.git_state()[source]¶
Git state of the build if known, otherwise empty. State is “CLEAN” if the repository was not changed from
git_commit(), “DIRTY” otherwise.
espressomd.visualization module¶
- class espressomd.visualization.Camera(cam_pos=array([0, 0, 1]), cam_target=array([0, 0, 0]), cam_right=array([1., 0., 0.]), move_speed=0.5, global_rot_speed=3.0, center=array([0, 0, 0]))[source]¶
Bases:
object
- class espressomd.visualization.Cylinder(shape, particle_type, color, material, quality, box_l, rasterize_resolution, rasterize_pointsize)[source]¶
Bases:
ShapeDrawable Shape Cylinder.
- class espressomd.visualization.Ellipsoid(shape, particle_type, color, material, quality, box_l, rasterize_resolution, rasterize_pointsize)[source]¶
Bases:
ShapeDrawable Shape Ellipsoid.
- class espressomd.visualization.HollowConicalFrustum(shape, particle_type, color, material, quality, box_l, rasterize_resolution, rasterize_pointsize)[source]¶
Bases:
ShapeDrawable Shape HollowConicalFrustum.
- class espressomd.visualization.KeyboardButtonEvent(button, fireEvent, callback, internal=False)[source]¶
Bases:
objectKeyboard event used for keyboard callbacks. Stores button, event type and callback.
- class espressomd.visualization.KeyboardFireEvent[source]¶
Bases:
objectEvent type of button used for keyboard callbacks.
- Hold = 1¶
- Pressed = 0¶
- Released = 2¶
- class espressomd.visualization.MouseButtonEvent(button, fireEvent, callback, positional=False)[source]¶
Bases:
objectMouse event used for mouse callbacks. Stores button and callback.
- class espressomd.visualization.MouseFireEvent[source]¶
Bases:
objectEvent type of mouse button used for mouse callbacks.
- ButtonMotion = 2¶
- ButtonPressed = 0¶
- ButtonReleased = 3¶
- DoubleClick = 4¶
- FreeMotion = 1¶
- class espressomd.visualization.Shape(shape, particle_type, color, material, quality, box_l, rasterize_resolution, rasterize_pointsize)[source]¶
Bases:
objectShape base class in the visualizer context.
- class espressomd.visualization.SimplePore(shape, particle_type, color, material, quality, box_l, rasterize_resolution, rasterize_pointsize)[source]¶
Bases:
ShapeDrawable Shape SimplePore.
- class espressomd.visualization.Slitpore(shape, particle_type, color, material, quality, box_l, rasterize_resolution, rasterize_pointsize)[source]¶
Bases:
ShapeDrawable Shape Slitpore.
- class espressomd.visualization.Sphere(shape, particle_type, color, material, quality, box_l, rasterize_resolution, rasterize_pointsize)[source]¶
Bases:
ShapeDrawable Shape Sphere.
- class espressomd.visualization.Spherocylinder(shape, particle_type, color, material, quality, box_l, rasterize_resolution, rasterize_pointsize)[source]¶
Bases:
ShapeDrawable Shape Spherocylinder.
- class espressomd.visualization.Wall(shape, particle_type, color, material, quality, box_l, rasterize_resolution, rasterize_pointsize)[source]¶
Bases:
ShapeDrawable Shape Wall.
- espressomd.visualization.draw_cylinder(posA, posB, radius, color, material, quality, draw_caps=False)[source]¶
- class espressomd.visualization.openGLLive(system, **kwargs)[source]¶
Bases:
objectThis class provides live visualization using pyOpenGL. Use the update method to push your current simulation state after integrating. Modify the appearance with a list of keywords. Timed callbacks can be registered via the
register_callback()method and keyboard callbacks viakeyboard_manager.register_button().- Parameters:
system (
espressomd.system.System)window_size ((2,) array_like of
int, optional) – Size of the visualizer window in pixels.name (
str, optional) – The name of the visualizer window.background_color ((3,) array_like of
float, optional) – RGB of the background.periodic_images ((3,) array_like of
int, optional) – Periodic repetitions on both sides of the box in xyz-direction.draw_box (
bool, optional) – Draw wireframe boundaries.draw_axis (
bool, optional) – Draw xyz system axes.draw_nodes (
bool, optional) – Draw node boxes.draw_cells (
bool, optional) – Draw cell boxes.quality_particles (
int, optional) – The number of subdivisions for particle spheres.quality_bonds (
int, optional) – The number of subdivisions for cylindrical bonds.quality_arrows (
int, optional) – The number of subdivisions for external force arrows.quality_constraints (
int, optional) – The number of subdivisions for primitive constraints.close_cut_distance (
float, optional) – The distance from the viewer to the near clipping plane.far_cut_distance (
float, optional) – The distance from the viewer to the far clipping plane.camera_position (
stror (3,) array_like offloat, optional) – Initial camera position. Use'auto'(default) for shifted position in z-direction.camera_target (
stror (3,) array_like offloat, optional) – Initial camera target. Use'auto'(default) to look towards the system center.camera_right ((3,) array_like of
float, optional) – Camera right vector in system coordinates. Default is[1, 0, 0]particle_sizes (
stror array_like offloator callable, optional) –'auto'(default): The Lennard-Jones sigma value of the self-interaction is used for the particle diameter.callable: A lambda function with one argument. Internally, the numerical particle type is passed to the lambda function to determine the particle radius.
list: A list of particle radii, indexed by the particle type.
particle_coloring (
str, optional) –'auto'(default): Colors of charged particles are specified byparticle_charge_colors, neutral particles byparticle_type_colors.'charge': Minimum and maximum charge of all particles is determined by the visualizer. All particles are colored by a linear interpolation of the two colors given byparticle_charge_colorsaccording to their charge.'type': Particle colors are specified byparticle_type_colors, indexed by their numerical particle type.'node': Color according to the node the particle is on.
particle_type_colors (array_like of
float, optional) – Colors for particle types.particle_type_materials (array_like of
str, optional) – Materials of the particle types.particle_charge_colors ((2,) array_like of
float, optional) – Two colors for min/max charged particles.draw_constraints (
bool, optional) – Enables constraint visualization. For simple constraints (planes, spheres and cylinders), OpenGL primitives are used. Otherwise, visualization by rasterization is used.rasterize_pointsize (
float, optional) – Point size for the rasterization dots.rasterize_resolution (
float, optional) – Accuracy of the rasterization.quality_constraints (
int, optional) – The number of subdivisions for primitive constraints.constraint_type_colors (array_like of
float, optional) – Colors of the constraints by type.constraint_type_materials (array_like of
str, optional) – Materials of the constraints by type.draw_bonds (
bool, optional) – Enables bond visualization.bond_type_radius (array_like of
float, optional) – Radii of bonds by type.bond_type_colors (array_like of
float, optional) – Color of bonds by type.bond_type_materials (array_like of
str, optional) – Materials of bonds by type.ext_force_arrows (
bool, optional) – Enables external force visualization.ext_force_arrows_type_scale (array_like of
float, optional) – List of scale factors of external force arrows for different particle types.ext_force_arrows_type_colors (array_like of
float, optional) – Colors of ext_force arrows for different particle types.ext_force_arrows_type_materials (array_like of
str, optional) – Materials of ext_force arrows for different particle types.ext_force_arrows_type_radii (array_like of
float, optional) – List of arrow radii for different particle types.force_arrows (
bool, optional) – Enables particle force visualization.force_arrows_type_scale (array_like of
float, optional) – List of scale factors of particle force arrows for different particle types.force_arrows_type_colors (array_like of
float, optional) – Colors of particle force arrows for different particle types.force_arrows_type_materials (array_like of
str, optional) – Materials of particle force arrows for different particle types.force_arrows_type_radii (array_like of
float, optional) – List of arrow radii for different particle types.velocity_arrows (
bool, optional) – Enables particle velocity visualization.velocity_arrows_type_scale (array_like of
float, optional) – List of scale factors of particle velocity arrows for different particle types.velocity_arrows_type_colors (array_like of
float, optional) – Colors of particle velocity arrows for different particle types.velocity_arrows_type_materials (array_like of
str, optional) – Materials of particle velocity arrows for different particle types.velocity_arrows_type_radii (array_like of
float, optional) – List of arrow radii for different particle types.director_arrows (
bool, optional) – Enables particle director visualization.director_arrows_type_scale (
float, optional) – Scale factor of particle director arrows for different particle types.director_arrows_type_colors (array_like of
float, optional) – Colors of particle director arrows for different particle types.director_arrows_type_materials (array_like of
str, optional) – Materials of particle director arrows for different particle types.director_arrows_type_radii (array_like of
float, optional) – List of arrow radii for different particle types.drag_enabled (
bool, optional) – Enables mouse-controlled particles dragging (Default: False)drag_force (
bool, optional) – Factor for particle draggingLB_draw_nodes (
bool, optional) – Draws a lattice representation of the LB nodes that are no boundaries.LB_draw_node_boundaries (
bool, optional) – Draws a lattice representation of the LB nodes that are boundaries.LB_draw_boundaries (
bool, optional) – Draws the LB shapes.LB_draw_velocity_plane (
bool, optional) – Draws LB node velocity arrows in a plane perpendicular to the axis inLB_plane_axis, at a distanceLB_plane_distfrom the origin, everyLB_plane_ngridgrid points.LB_plane_axis (
int, optional) – LB node velocity arrows are drawn in a plane perpendicular to the x, y, or z axes, which are encoded by values 0, 1, or 2 respectively.LB_plane_dist (
float, optional) – LB node velocity arrows are drawn in a plane perpendicular to the x, y, or z axes, at a distanceLB_plane_distfrom the origin.LB_plane_ngrid (
int, optional) – LB node velocity arrows are drawn in a plane perpendicular to the x, y, or z axes, everyLB_plane_ngridgrid points.LB_vel_scale (
float, optional) – Rescale LB node velocity arrow length.LB_vel_radius_scale (
float, optional) – Rescale LB node velocity arrow radii.LB_arrow_color_fluid ((3,) array_like of
float, optional) – RGB of the LB velocity arrows inside the fluid.LB_arrow_color_boundary ((3,) array_like of
float, optional) – RGB of the LB velocity arrows inside boundaries.LB_arrow_material (
str, optional) – Material of LB arrows.quality_constraints (
int, optional) – The number of subdivisions for LB arrows.light_pos ((3,) array_like of
float, optional) – If auto (default) is used, the light is placed dynamically in the particle barycenter of the system. Otherwise, a fixed coordinate can be set.light_colors (array_like of
float, optional) – Three lists to specify ambient, diffuse and specular light colors.light_brightness (
float, optional) – Brightness (inverse constant attenuation) of the light.light_size (
float, optional) – Size (inverse linear attenuation) of the light. If auto (default) is used, the light size will be set to a reasonable value according to the box size at start.spotlight_enabled (
bool, optional) – If set to True (default), it enables a spotlight on the camera position pointing in look direction.spotlight_colors (array_like of
float, optional) – Three lists to specify ambient, diffuse and specular spotlight colors.spotlight_angle (
float, optional) – The spread angle of the spotlight in degrees (from 0 to 90).spotlight_brightness (
float, optional) – Brightness (inverse constant attenuation) of the spotlight.spotlight_focus (
float, optional) – Focus (spot exponent) for the spotlight from 0 (uniform) to 128.
Notes
The visualization of some constraints is either improved by or even relies on the presence of an installed OpenGL Extrusion library on your system.
- materials = {'bright': [0.9, 1.0, 0.8, 0.4, 1.0], 'chrome': [0.25, 0.4, 0.774597, 0.6, 1.0], 'dark': [0.4, 0.5, 0.1, 0.4, 1.0], 'medium': [0.6, 0.8, 0.2, 0.4, 1.0], 'plastic': [0, 0.55, 0.7, 0.25, 1.0], 'rubber': [0, 0.4, 0.7, 0.078125, 1.0], 'steel': [0.25, 0.38, 0, 0.32, 1.0], 'transparent1': [0.6, 0.8, 0.2, 0.5, 0.8], 'transparent2': [0.6, 0.8, 0.2, 0.5, 0.4], 'transparent3': [0.6, 0.8, 0.2, 0.5, 0.2]}¶
- screenshot(path)[source]¶
Renders the current state into an image file at
pathwith dimensions ofspecs['window_size']in PNG format.